LKCell: Efficient Cell Nuclei Instance Segmentation with Large Convolution Kernels
Paper and Code
Jul 25, 2024


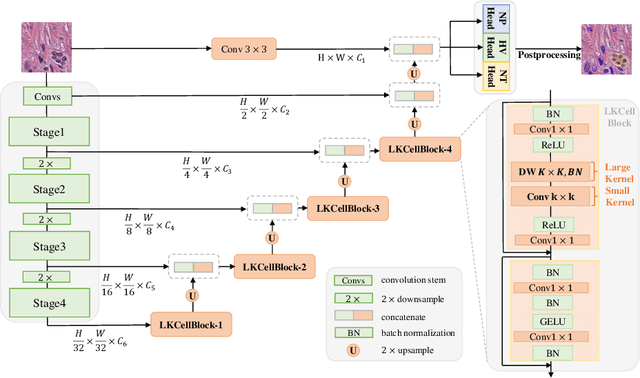
The segmentation of cell nuclei in tissue images stained with the blood dye hematoxylin and eosin (H$\&$E) is essential for various clinical applications and analyses. Due to the complex characteristics of cellular morphology, a large receptive field is considered crucial for generating high-quality segmentation. However, previous methods face challenges in achieving a balance between the receptive field and computational burden. To address this issue, we propose LKCell, a high-accuracy and efficient cell segmentation method. Its core insight lies in unleashing the potential of large convolution kernels to achieve computationally efficient large receptive fields. Specifically, (1) We transfer pre-trained large convolution kernel models to the medical domain for the first time, demonstrating their effectiveness in cell segmentation. (2) We analyze the redundancy of previous methods and design a new segmentation decoder based on large convolution kernels. It achieves higher performance while significantly reducing the number of parameters. We evaluate our method on the most challenging benchmark and achieve state-of-the-art results (0.5080 mPQ) in cell nuclei instance segmentation with only 21.6% FLOPs compared with the previous leading method. Our source code and models are available at https://github.com/hustvl/LKCell.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge