Jeya Balasubramanian
TMA-Grid: An open-source, zero-footprint web application for FAIR Tissue MicroArray De-arraying
Jul 30, 2024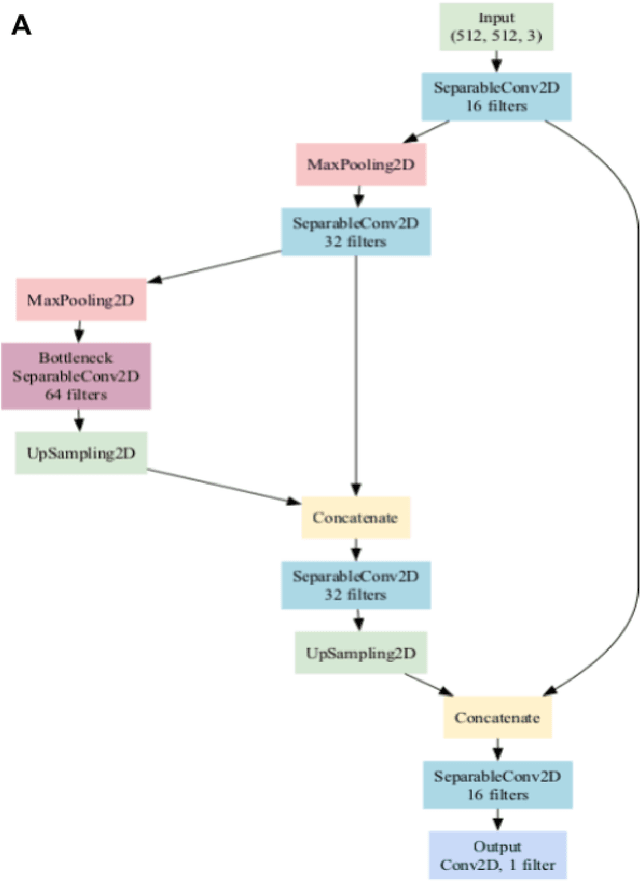
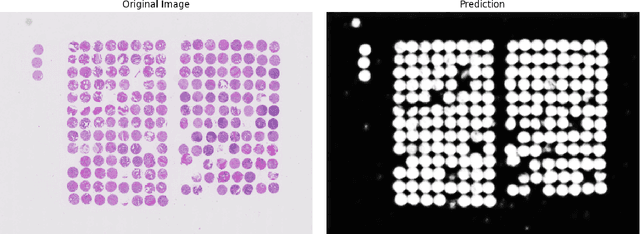


Abstract:Background: Tissue Microarrays (TMAs) significantly increase analytical efficiency in histopathology and large-scale epidemiologic studies by allowing multiple tissue cores to be scanned on a single slide. The individual cores can be digitally extracted and then linked to metadata for analysis in a process known as de-arraying. However, TMAs often contain core misalignments and artifacts due to assembly errors, which can adversely affect the reliability of the extracted cores during the de-arraying process. Moreover, conventional approaches for TMA de-arraying rely on desktop solutions.Therefore, a robust yet flexible de-arraying method is crucial to account for these inaccuracies and ensure effective downstream analyses. Results: We developed TMA-Grid, an in-browser, zero-footprint, interactive web application for TMA de-arraying. This web application integrates a convolutional neural network for precise tissue segmentation and a grid estimation algorithm to match each identified core to its expected location. The application emphasizes interactivity, allowing users to easily adjust segmentation and gridding results. Operating entirely in the web-browser, TMA-Grid eliminates the need for downloads or installations and ensures data privacy. Adhering to FAIR principles (Findable, Accessible, Interoperable, and Reusable), the application and its components are designed for seamless integration into TMA research workflows. Conclusions: TMA-Grid provides a robust, user-friendly solution for TMA dearraying on the web. As an open, freely accessible platform, it lays the foundation for collaborative analyses of TMAs and similar histopathology imaging data. Availability: Web application: https://episphere.github.io/tma-grid Code: https://github.com/episphere/tma-grid Tutorial: https://youtu.be/miajqyw4BVk
FastImpute: A Baseline for Open-source, Reference-Free Genotype Imputation Methods -- A Case Study in PRS313
Jul 12, 2024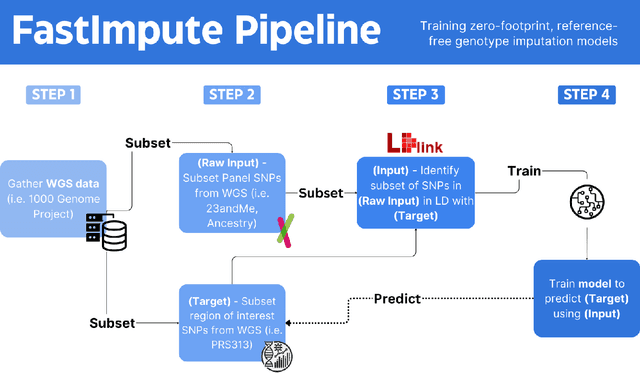
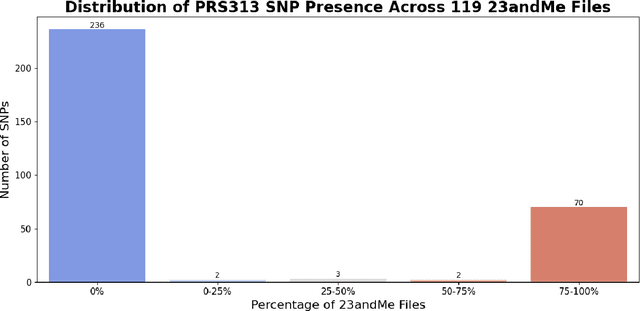
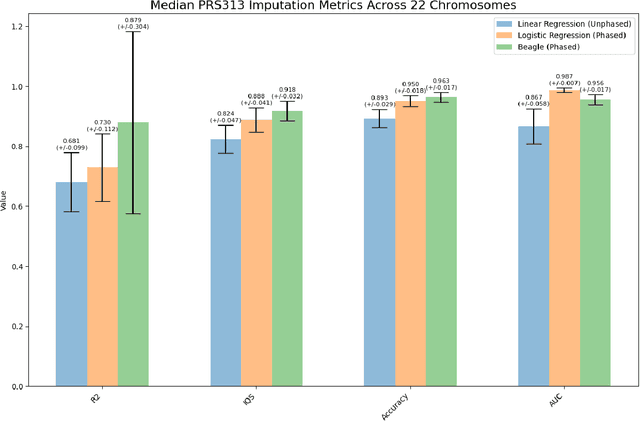
Abstract:Genotype imputation enhances genetic data by predicting missing SNPs using reference haplotype information. Traditional methods leverage linkage disequilibrium (LD) to infer untyped SNP genotypes, relying on the similarity of LD structures between genotyped target sets and fully sequenced reference panels. Recently, reference-free deep learning-based methods have emerged, offering a promising alternative by predicting missing genotypes without external databases, thereby enhancing privacy and accessibility. However, these methods often produce models with tens of millions of parameters, leading to challenges such as the need for substantial computational resources to train and inefficiency for client-sided deployment. Our study addresses these limitations by introducing a baseline for a novel genotype imputation pipeline that supports client-sided imputation models generalizable across any genotyping chip and genomic region. This approach enhances patient privacy by performing imputation directly on edge devices. As a case study, we focus on PRS313, a polygenic risk score comprising 313 SNPs used for breast cancer risk prediction. Utilizing consumer genetic panels such as 23andMe, our model democratizes access to personalized genetic insights by allowing 23andMe users to obtain their PRS313 score. We demonstrate that simple linear regression can significantly improve the accuracy of PRS313 scores when calculated using SNPs imputed from consumer gene panels, such as 23andMe. Our linear regression model achieved an R^2 of 0.86, compared to 0.33 without imputation and 0.28 with simple imputation (substituting missing SNPs with the minor allele frequency). These findings suggest that popular SNP analysis libraries could benefit from integrating linear regression models for genotype imputation, providing a viable and light-weight alternative to reference based imputation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge