Javier Pérez de Frutos
AI-Dentify: Deep learning for proximal caries detection on bitewing x-ray -- HUNT4 Oral Health Study
Sep 30, 2023Abstract:Background: Dental caries diagnosis requires the manual inspection of diagnostic bitewing images of the patient, followed by a visual inspection and probing of the identified dental pieces with potential lesions. Yet the use of artificial intelligence, and in particular deep-learning, has the potential to aid in the diagnosis by providing a quick and informative analysis of the bitewing images. Methods: A dataset of 13,887 bitewings from the HUNT4 Oral Health Study were annotated individually by six different experts, and used to train three different object detection deep-learning architectures: RetinaNet (ResNet50), YOLOv5 (M size), and EfficientDet (D0 and D1 sizes). A consensus dataset of 197 images, annotated jointly by the same six dentist, was used for evaluation. A five-fold cross validation scheme was used to evaluate the performance of the AI models. Results: the trained models show an increase in average precision and F1-score, and decrease of false negative rate, with respect to the dental clinicians. Out of the three architectures studied, YOLOv5 shows the largest improvement, reporting 0.647 mean average precision, 0.548 mean F1-score, and 0.149 mean false negative rate. Whereas the best annotators on each of these metrics reported 0.299, 0.495, and 0.164 respectively. Conclusion: Deep-learning models have shown the potential to assist dental professionals in the diagnosis of caries. Yet, the task remains challenging due to the artifacts natural to the bitewings.
Train smarter, not harder: learning deep abdominal CT registration on scarce data
Nov 30, 2022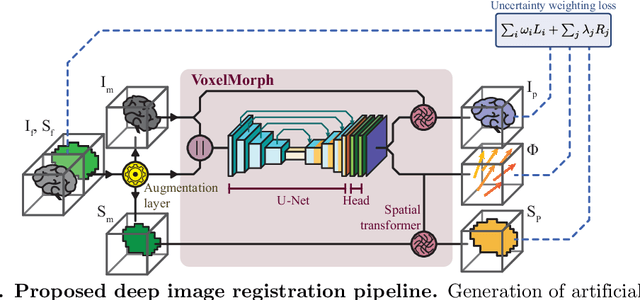
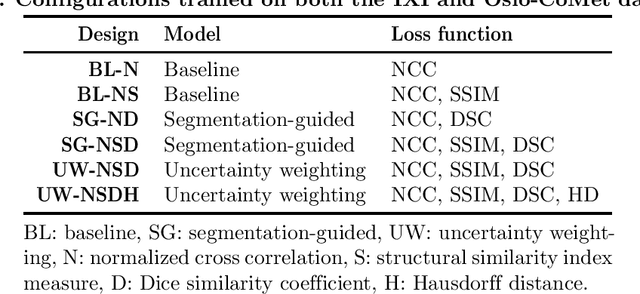
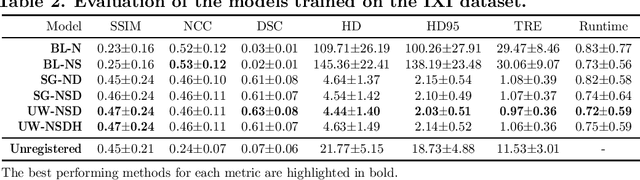
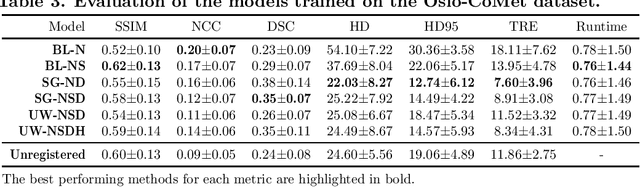
Abstract:Purpose: This study aims to explore training strategies to improve convolutional neural network-based image-to-image registration for abdominal imaging. Methods: Different training strategies, loss functions, and transfer learning schemes were considered. Furthermore, an augmentation layer which generates artificial training image pairs on-the-fly was proposed, in addition to a loss layer that enables dynamic loss weighting. Results: Guiding registration using segmentations in the training step proved beneficial for deep-learning-based image registration. Finetuning the pretrained model from the brain MRI dataset to the abdominal CT dataset further improved performance on the latter application, removing the need for a large dataset to yield satisfactory performance. Dynamic loss weighting also marginally improved performance, all without impacting inference runtime. Conclusion: Using simple concepts, we improved the performance of a commonly used deep image registration architecture, VoxelMorph. In future work, our framework, DDMR, should be validated on different datasets to further assess its value.
FastPathology: An open-source platform for deep learning-based research and decision support in digital pathology
Nov 11, 2020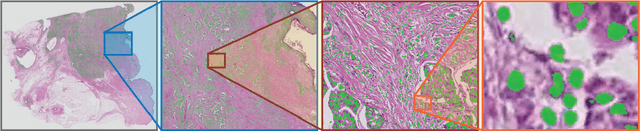
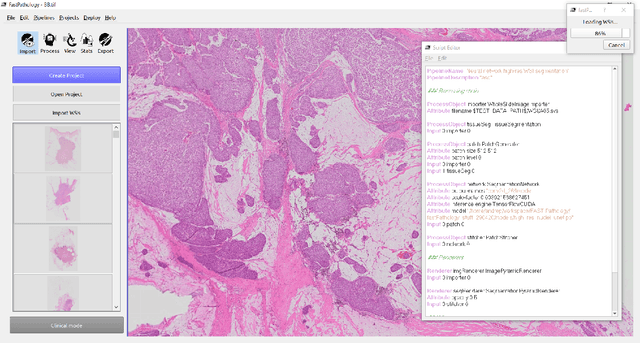
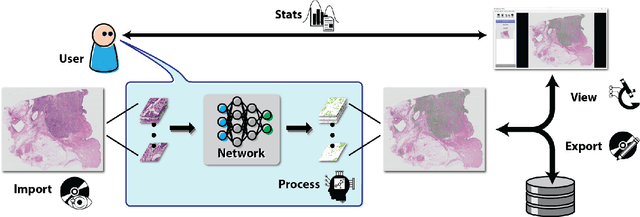
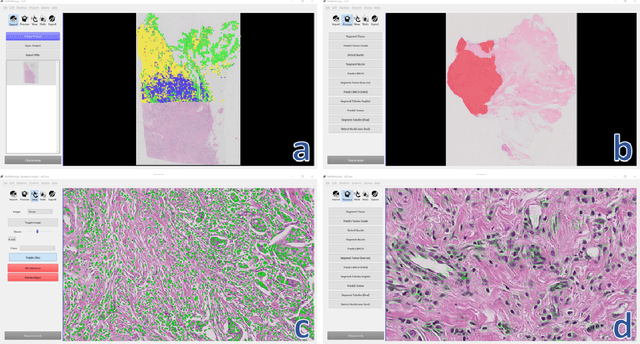
Abstract:Deep convolutional neural networks (CNNs) are the current state-of-the-art for digital analysis of histopathological images. The large size of whole-slide microscopy images (WSIs) requires advanced memory handling to read, display and process these images. There are several open-source platforms for working with WSIs, but few support deployment of CNN models. These applications use third-party solutions for inference, making them less user-friendly and unsuitable for high-performance image analysis. To make deployment of CNNs user-friendly and feasible on low-end machines, we have developed a new platform, FastPathology, using the FAST framework and C++. It minimizes memory usage for reading and processing WSIs, deployment of CNN models, and real-time interactive visualization of results. Runtime experiments were conducted on four different use cases, using different architectures, inference engines, hardware configurations and operating systems. Memory usage for reading, visualizing, zooming and panning a WSI were measured, using FastPathology and three existing platforms. FastPathology performed similarly in terms of memory to the other C++ based application, while using considerably less than the two Java-based platforms. The choice of neural network model, inference engine, hardware and processors influenced runtime considerably. Thus, FastPathology includes all steps needed for efficient visualization and processing of WSIs in a single application, including inference of CNNs with real-time display of the results. Source code, binary releases and test data can be found online on GitHub at https://github.com/SINTEFMedtek/FAST-Pathology/.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge