Jacob Antunes
Spatial-And-Context aware (SpACe) "virtual biopsy" radiogenomic maps to target tumor mutational status on structural MRI
Jun 17, 2020
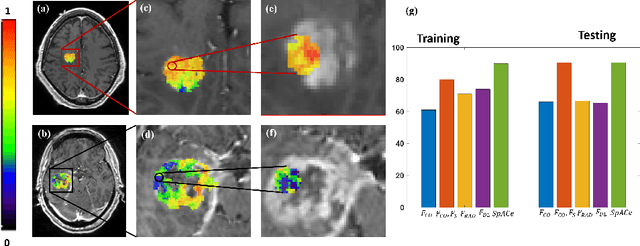
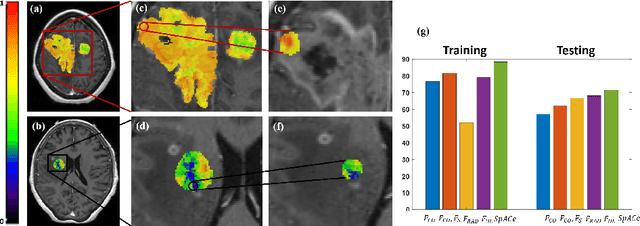
Abstract:With growing emphasis on personalized cancer-therapies,radiogenomics has shown promise in identifying target tumor mutational status on routine imaging (i.e. MRI) scans. These approaches fall into 2 categories: (1) deep-learning/radiomics (context-based), using image features from the entire tumor to identify the gene mutation status, or (2) atlas (spatial)-based to obtain likelihood of gene mutation status based on population statistics. While many genes (i.e. EGFR, MGMT) are spatially variant, a significant challenge in reliable assessment of gene mutation status on imaging has been the lack of available co-localized ground truth for training the models. We present Spatial-And-Context aware (SpACe) "virtual biopsy" maps that incorporate context-features from co-localized biopsy site along with spatial-priors from population atlases, within a Least Absolute Shrinkage and Selection Operator (LASSO) regression model, to obtain a per-voxel probability of the presence of a mutation status (M+ vs M-). We then use probabilistic pair-wise Markov model to improve the voxel-wise prediction probability. We evaluate the efficacy of SpACe maps on MRI scans with co-localized ground truth obtained from corresponding biopsy, to predict the mutation status of 2 driver genes in Glioblastoma: (1) EGFR (n=91), and (2) MGMT (n=81). When compared against deep-learning (DL) and radiomic models, SpACe maps obtained training and testing accuracies of 90% (n=71) and 90.48% (n=21) in identifying EGFR amplification status,compared to 80% and 71.4% via radiomics, and 74.28% and 65.5% via DL. For MGMT status, training and testing accuracies using SpACe were 88.3% (n=61) and 71.5% (n=20), compared to 52.4% and 66.7% using radiomics,and 79.3% and 68.4% using DL. Following validation,SpACe maps could provide surgical navigation to improve localization of sampling sites for targeting of specific driver genes in cancer.
MRQy: An Open-Source Tool for Quality Control of MR Imaging Data
Apr 13, 2020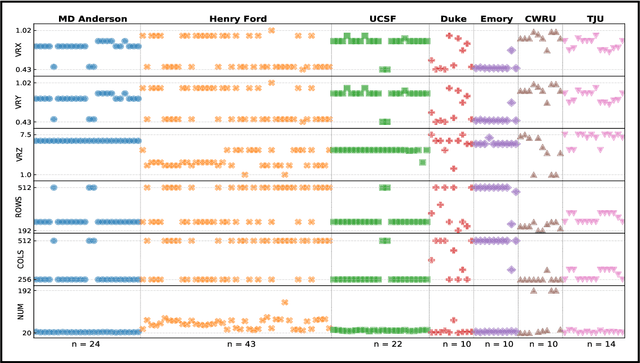
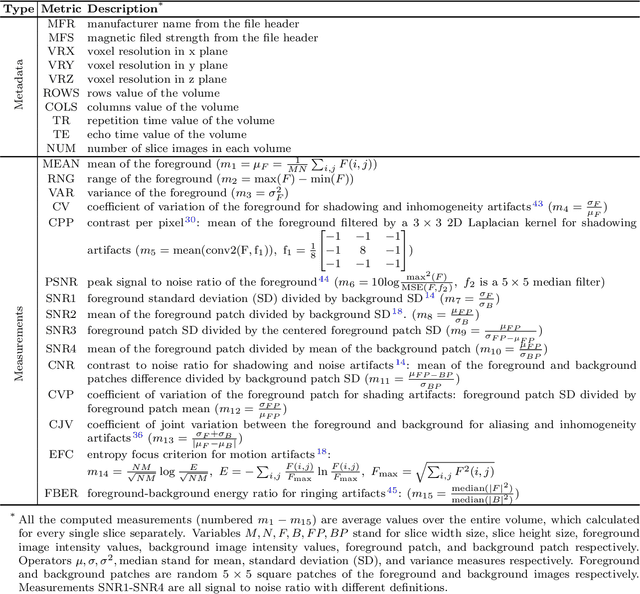

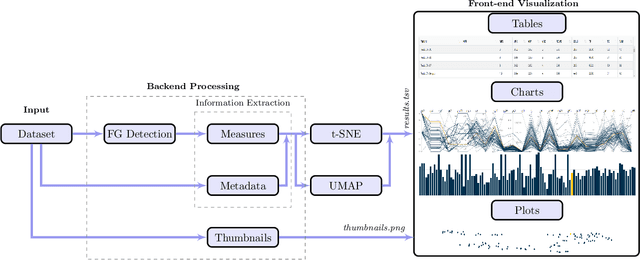
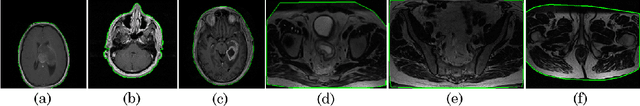
Abstract:Even as public data repositories such as The Cancer Imaging Archive (TCIA) have enabled development of new radiomics and machine learning schemes, a key concern remains the generalizability of these methods to unseen datasets. For MRI datasets, model performance could be impacted by (a) site- or scanner-specific variations in image resolution, field-of-view, or image contrast, or (b) presence of imaging artifacts such as noise, motion, inhomogeneity, ringing, or aliasing; which can adversely affect relative image quality between data cohorts. This indicates a need for a quantitative tool to quickly determine relative differences in MRI volumes both within and between large data cohorts. We present MRQy, a new open-source quality control tool to (a) interrogate MRI cohorts for site- or equipment-based differences, and (b) quantify the impact of MRI artifacts on relative image quality; to help determine how to correct for these variations prior to model development. MRQy extracts a series of quality measures (e.g. noise ratios, variation metrics, entropy and energy criteria) and MR image metadata (e.g. voxel resolution, image dimensions) for subsequent interrogation via a specialized HTML5 based front-end designed for real-time filtering and trend visualization. MRQy is designed to be a standalone, unsupervised tool that can be efficiently run on a standard desktop computer. It has been made freely accessible at http://github.com/ccipd/MRQy for wider community use and feedback. MRQy was used to evaluate (a) n=133 brain MRIs from TCIA (7 sites), and (b) n=104 rectal MRIs (3 local sites). MRQy measures revealed significant site-specific variations in both cohorts, indicating potential batch effects. Marked differences in specific MRQy measures were also able to identify MRI datasets that needed to be corrected for common MR imaging artifacts.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge