Guillaume Bollot
ChAda-ViT : Channel Adaptive Attention for Joint Representation Learning of Heterogeneous Microscopy Images
Nov 26, 2023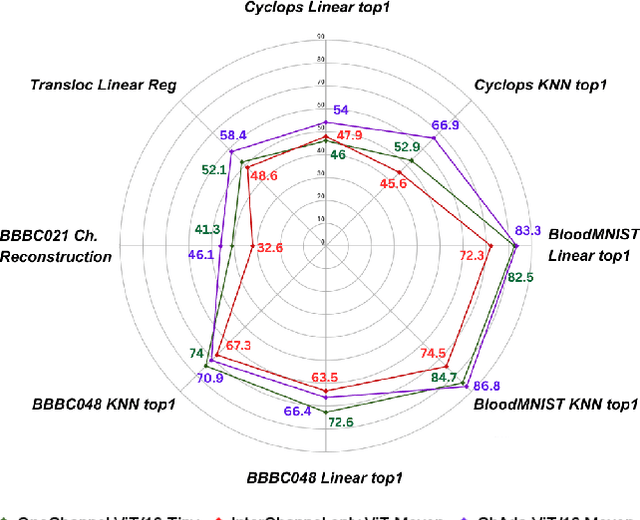
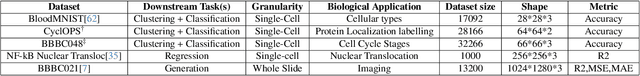
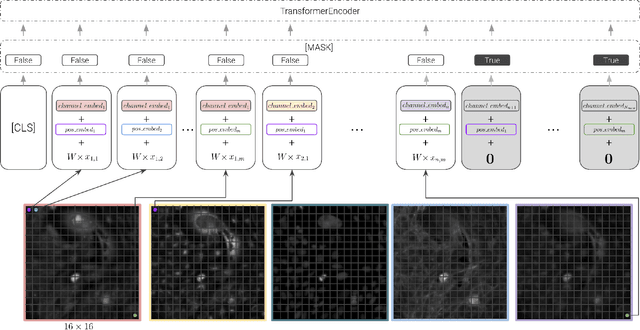

Abstract:Unlike color photography images, which are consistently encoded into RGB channels, biological images encompass various modalities, where the type of microscopy and the meaning of each channel varies with each experiment. Importantly, the number of channels can range from one to a dozen and their correlation is often comparatively much lower than RGB, as each of them brings specific information content. This aspect is largely overlooked by methods designed out of the bioimage field, and current solutions mostly focus on intra-channel spatial attention, often ignoring the relationship between channels, yet crucial in most biological applications. Importantly, the variable channel type and count prevent the projection of several experiments to a unified representation for large scale pre-training. In this study, we propose ChAda-ViT, a novel Channel Adaptive Vision Transformer architecture employing an Inter-Channel Attention mechanism on images with an arbitrary number, order and type of channels. We also introduce IDRCell100k, a bioimage dataset with a rich set of 79 experiments covering 7 microscope modalities, with a multitude of channel types, and channel counts varying from 1 to 10 per experiment. Our proposed architecture, trained in a self-supervised manner, outperforms existing approaches in several biologically relevant downstream tasks. Additionally, it can be used to bridge the gap for the first time between assays with different microscopes, channel numbers or types by embedding various image and experimental modalities into a unified biological image representation. The latter should facilitate interdisciplinary studies and pave the way for better adoption of deep learning in biological image-based analyses. Code and Data to be released soon.
No Free Lunch in Self Supervised Representation Learning
Apr 23, 2023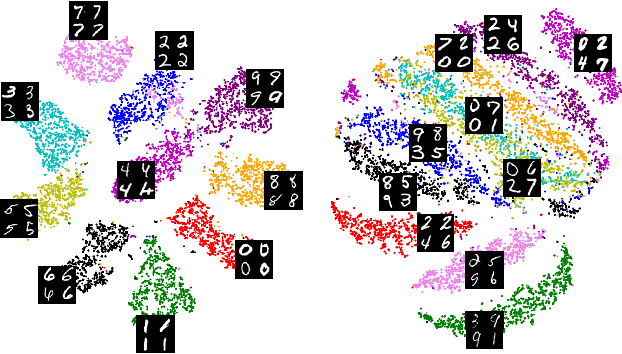

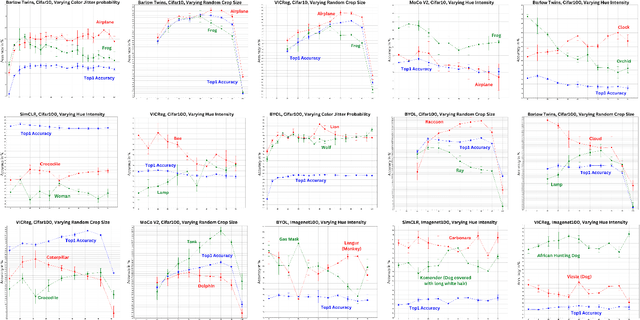
Abstract:Self-supervised representation learning in computer vision relies heavily on hand-crafted image transformations to learn meaningful and invariant features. However few extensive explorations of the impact of transformation design have been conducted in the literature. In particular, the dependence of downstream performances to transformation design has been established, but not studied in depth. In this work, we explore this relationship, its impact on a domain other than natural images, and show that designing the transformations can be viewed as a form of supervision. First, we demonstrate that not only do transformations have an effect on downstream performance and relevance of clustering, but also that each category in a supervised dataset can be impacted in a different way. Following this, we explore the impact of transformation design on microscopy images, a domain where the difference between classes is more subtle and fuzzy than in natural images. In this case, we observe a greater impact on downstream tasks performances. Finally, we demonstrate that transformation design can be leveraged as a form of supervision, as careful selection of these by a domain expert can lead to a drastic increase in performance on a given downstream task.
Comparison of semi-supervised learning methods for High Content Screening quality control
Aug 09, 2022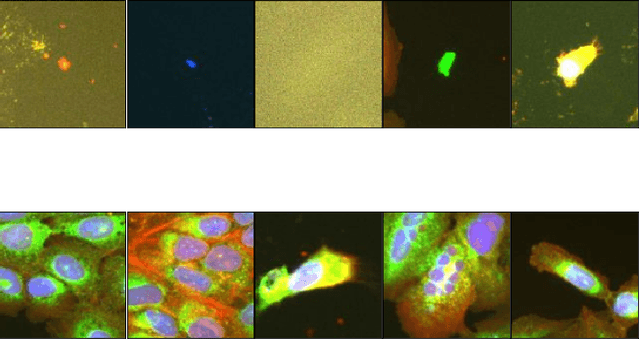

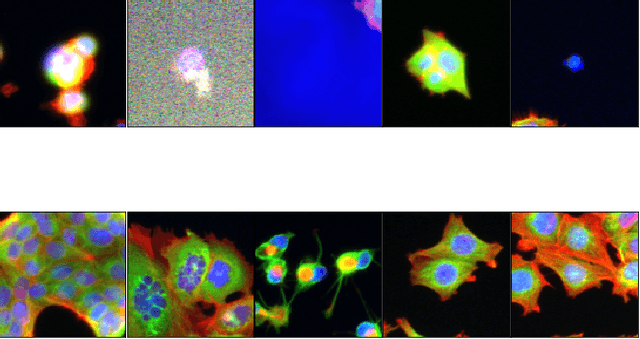
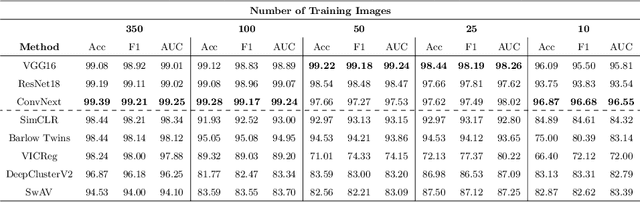
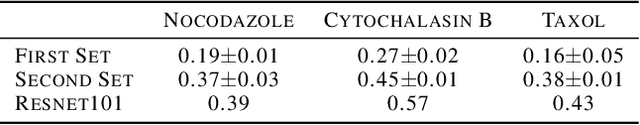
Abstract:Progress in automated microscopy and quantitative image analysis has promoted high-content screening (HCS) as an efficient drug discovery and research tool. While HCS offers to quantify complex cellular phenotypes from images at high throughput, this process can be obstructed by image aberrations such as out-of-focus image blur, fluorophore saturation, debris, a high level of noise, unexpected auto-fluorescence or empty images. While this issue has received moderate attention in the literature, overlooking these artefacts can seriously hamper downstream image processing tasks and hinder detection of subtle phenotypes. It is therefore of primary concern, and a prerequisite, to use quality control in HCS. In this work, we evaluate deep learning options that do not require extensive image annotations to provide a straightforward and easy to use semi-supervised learning solution to this issue. Concretely, we compared the efficacy of recent self-supervised and transfer learning approaches to provide a base encoder to a high throughput artefact image detector. The results of this study suggest that transfer learning methods should be preferred for this task as they not only performed best here but present the advantage of not requiring sensitive hyperparameter settings nor extensive additional training.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge