Goktug Guvercin
Conformable Convolution for Topologically Aware Learning of Complex Anatomical Structures
Dec 29, 2024Abstract:While conventional computer vision emphasizes pixel-level and feature-based objectives, medical image analysis of intricate biological structures necessitates explicit representation of their complex topological properties. Despite their successes, deep learning models often struggle to accurately capture the connectivity and continuity of fine, sometimes pixel-thin, yet critical structures due to their reliance on implicit learning from data. Such shortcomings can significantly impact the reliability of analysis results and hinder clinical decision-making. To address this challenge, we introduce Conformable Convolution, a novel convolutional layer designed to explicitly enforce topological consistency. Conformable Convolution learns adaptive kernel offsets that preferentially focus on regions of high topological significance within an image. This prioritization is guided by our proposed Topological Posterior Generator (TPG) module, which leverages persistent homology. The TPG module identifies key topological features and guides the convolutional layers by applying persistent homology to feature maps transformed into cubical complexes. Our proposed modules are architecture-agnostic, enabling them to be integrated seamlessly into various architectures. We showcase the effectiveness of our framework in the segmentation task, where preserving the interconnectedness of structures is critical. Experimental results on three diverse datasets demonstrate that our framework effectively preserves the topology in the segmentation downstream task, both quantitatively and qualitatively.
Machine Learning Methods for Brain Network Classification: Application to Autism Diagnosis using Cortical Morphological Networks
Apr 28, 2020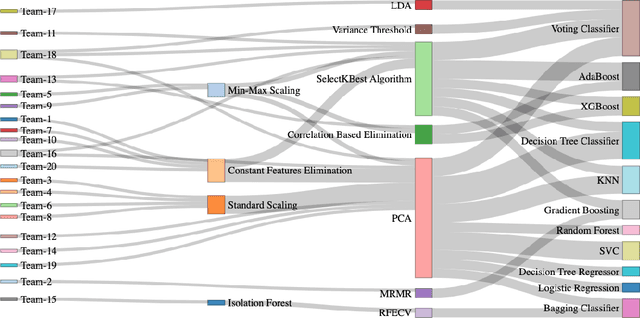
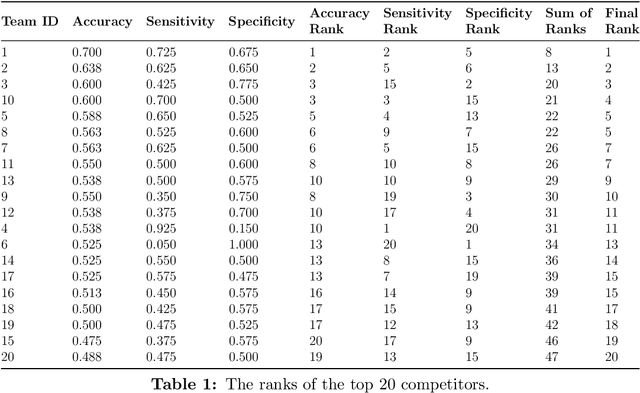
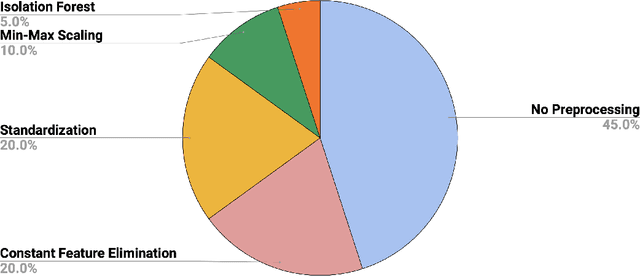
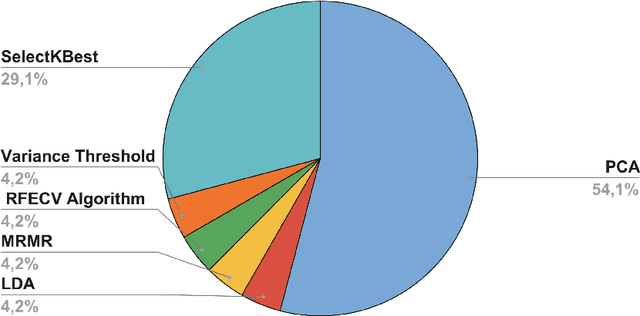
Abstract:Autism spectrum disorder (ASD) affects the brain connectivity at different levels. Nonetheless, non-invasively distinguishing such effects using magnetic resonance imaging (MRI) remains very challenging to machine learning diagnostic frameworks due to ASD heterogeneity. So far, existing network neuroscience works mainly focused on functional (derived from functional MRI) and structural (derived from diffusion MRI) brain connectivity, which might not capture relational morphological changes between brain regions. Indeed, machine learning (ML) studies for ASD diagnosis using morphological brain networks derived from conventional T1-weighted MRI are very scarce. To fill this gap, we leverage crowdsourcing by organizing a Kaggle competition to build a pool of machine learning pipelines for neurological disorder diagnosis with application to ASD diagnosis using cortical morphological networks derived from T1-weighted MRI. During the competition, participants were provided with a training dataset and only allowed to check their performance on a public test data. The final evaluation was performed on both public and hidden test datasets based on accuracy, sensitivity, and specificity metrics. Teams were ranked using each performance metric separately and the final ranking was determined based on the mean of all rankings. The first-ranked team achieved 70% accuracy, 72.5% sensitivity, and 67.5% specificity, while the second-ranked team achieved 63.8%, 62.5%, 65% respectively. Leveraging participants to design ML diagnostic methods within a competitive machine learning setting has allowed the exploration and benchmarking of wide spectrum of ML methods for ASD diagnosis using cortical morphological networks.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge