Gihan Panapitiya
FragNet: A Graph Neural Network for Molecular Property Prediction with Four Layers of Interpretability
Oct 16, 2024Abstract:Molecular property prediction is a crucial step in many modern-day scientific applications including drug discovery and energy storage material design. Despite the availability of numerous machine learning models for this task, we are lacking in models that provide both high accuracies and interpretability of the predictions. We introduce the FragNet architecture, a graph neural network not only capable of achieving prediction accuracies comparable to the current state-of-the-art models, but also able to provide insight on four levels of molecular substructures. This model enables understanding of which atoms, bonds, molecular fragments, and molecular fragment connections are critical in the prediction of a given molecular property. The ability to interpret the importance of connections between fragments is of particular interest for molecules which have substructures that are not connected with regular covalent bonds. The interpretable capabilities of FragNet are key to gaining scientific insights from the model's learned patterns between molecular structure and molecular properties.
Dynamic Molecular Graph-based Implementation for Biophysical Properties Prediction
Dec 20, 2022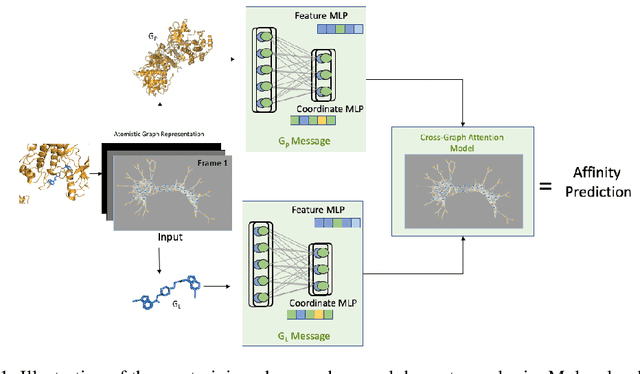
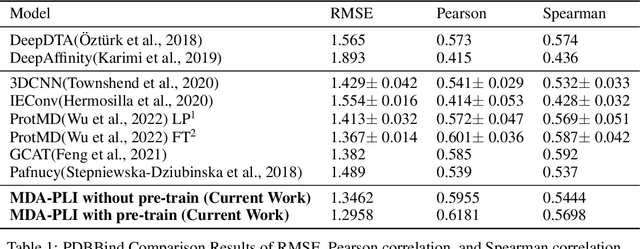
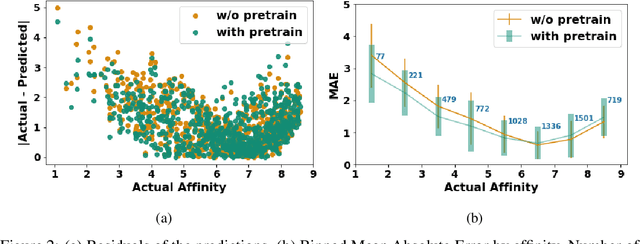
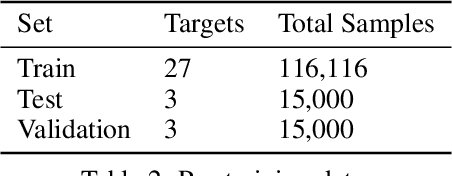
Abstract:Neural Networks (GNNs) have revolutionized the molecular discovery to understand patterns and identify unknown features that can aid in predicting biophysical properties and protein-ligand interactions. However, current models typically rely on 2-dimensional molecular representations as input, and while utilization of 2\3- dimensional structural data has gained deserved traction in recent years as many of these models are still limited to static graph representations. We propose a novel approach based on the transformer model utilizing GNNs for characterizing dynamic features of protein-ligand interactions. Our message passing transformer pre-trains on a set of molecular dynamic data based off of physics-based simulations to learn coordinate construction and make binding probability and affinity predictions as a downstream task. Through extensive testing we compare our results with the existing models, our MDA-PLI model was able to outperform the molecular interaction prediction models with an RMSE of 1.2958. The geometric encodings enabled by our transformer architecture and the addition of time series data add a new dimensionality to this form of research.
Predicting Aqueous Solubility of Organic Molecules Using Deep Learning Models with Varied Molecular Representations
May 27, 2021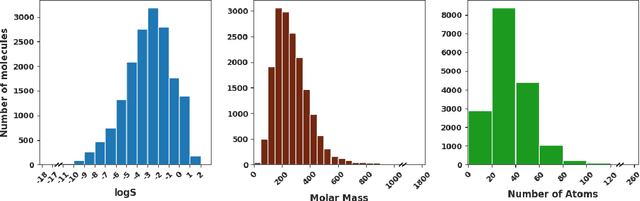
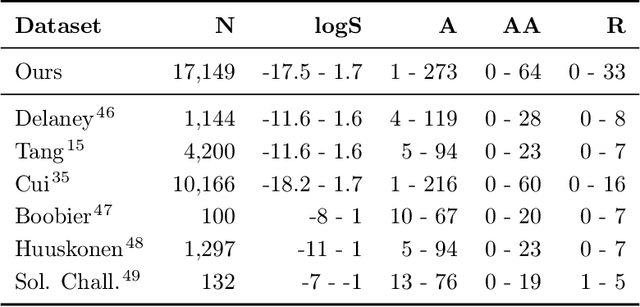
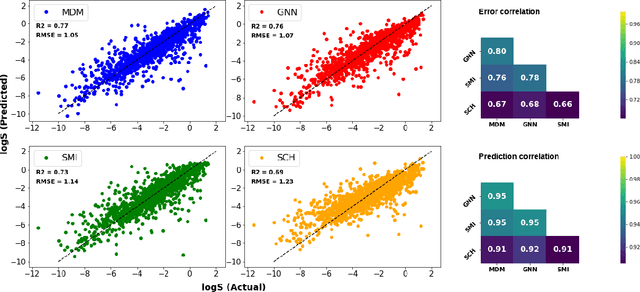
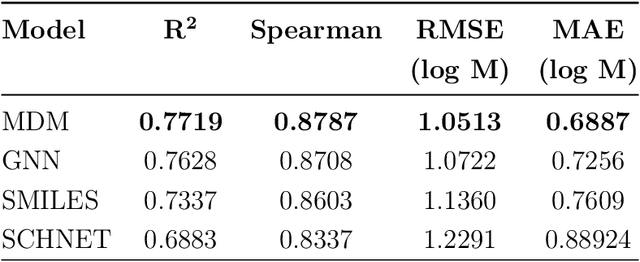
Abstract:Determining the aqueous solubility of molecules is a vital step in many pharmaceutical, environmental, and energy storage applications. Despite efforts made over decades, there are still challenges associated with developing a solubility prediction model with satisfactory accuracy for many of these applications. The goal of this study is to develop a general model capable of predicting the solubility of a broad range of organic molecules. Using the largest currently available solubility dataset, we implement deep learning-based models to predict solubility from molecular structure and explore several different molecular representations including molecular descriptors, simplified molecular-input line-entry system (SMILES) strings, molecular graphs, and three-dimensional (3D) atomic coordinates using four different neural network architectures - fully connected neural networks (FCNNs), recurrent neural networks (RNNs), graph neural networks (GNNs), and SchNet. We find that models using molecular descriptors achieve the best performance, with GNN models also achieving good performance. We perform extensive error analysis to understand the molecular properties that influence model performance, perform feature analysis to understand which information about molecular structure is most valuable for prediction, and perform a transfer learning and data size study to understand the impact of data availability on model performance.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge