George Heintz
MedML: Fusing Medical Knowledge and Machine Learning Models for Early Pediatric COVID-19 Hospitalization and Severity Prediction
Jul 25, 2022


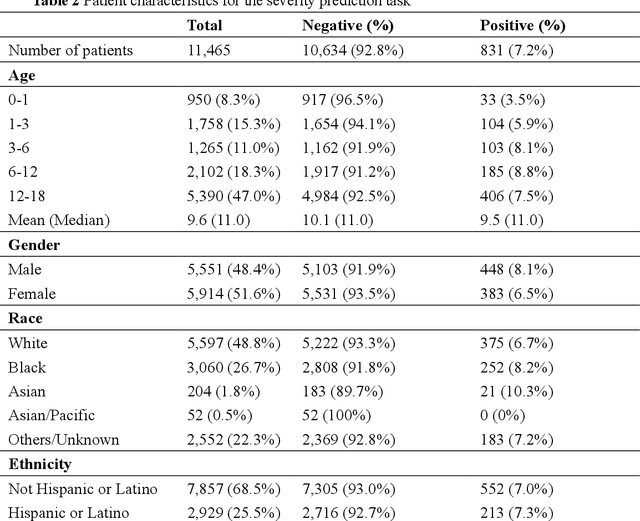
Abstract:The COVID-19 pandemic has caused devastating economic and social disruption, straining the resources of healthcare institutions worldwide. This has led to a nationwide call for models to predict hospitalization and severe illness in patients with COVID-19 to inform distribution of limited healthcare resources. We respond to one of these calls specific to the pediatric population. To address this challenge, we study two prediction tasks for the pediatric population using electronic health records: 1) predicting which children are more likely to be hospitalized, and 2) among hospitalized children, which individuals are more likely to develop severe symptoms. We respond to the national Pediatric COVID-19 data challenge with a novel machine learning model, MedML. MedML extracts the most predictive features based on medical knowledge and propensity scores from over 6 million medical concepts and incorporates the inter-feature relationships between heterogeneous medical features via graph neural networks (GNN). We evaluate MedML across 143,605 patients for the hospitalization prediction task and 11,465 patients for the severity prediction task using data from the National Cohort Collaborative (N3C) dataset. We also report detailed group-level and individual-level feature importance analyses to evaluate the model interpretability. MedML achieves up to a 7% higher AUROC score and up to a 14% higher AUPRC score compared to the best baseline machine learning models and performs well across all nine national geographic regions and over all three-month spans since the start of the pandemic. Our cross-disciplinary research team has developed a method of incorporating clinical domain knowledge as the framework for a new type of machine learning model that is more predictive and explainable than current state-of-the-art data-driven feature selection methods.
Towards a Comprehensive Solution for a Vision-based Digitized Neurological Examination
May 15, 2022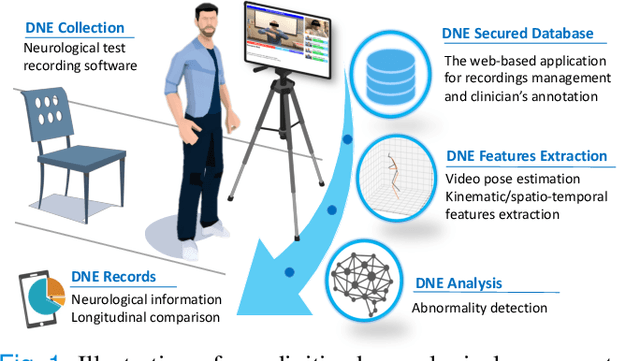
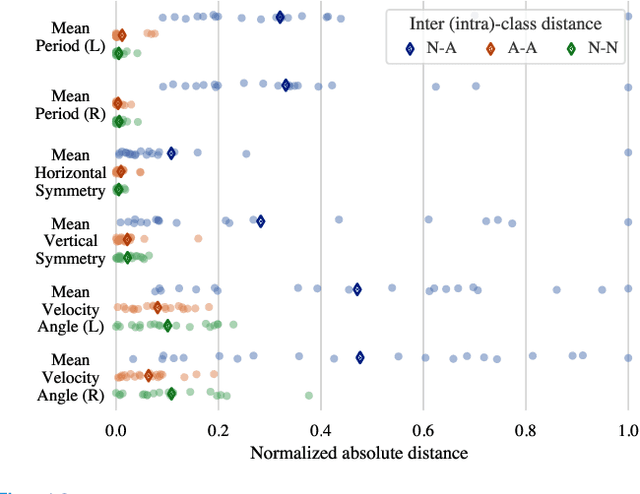
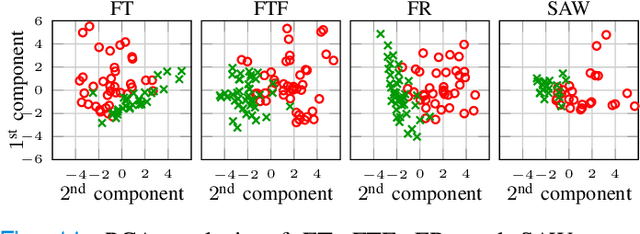
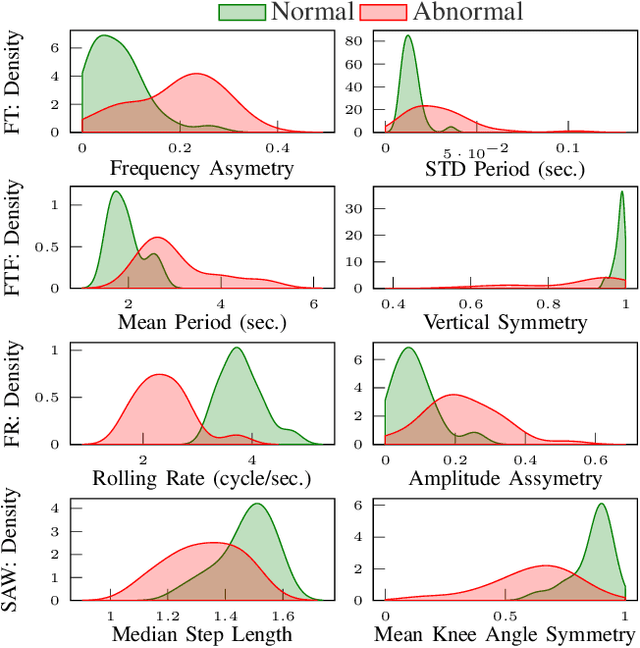
Abstract:The ability to use digitally recorded and quantified neurological exam information is important to help healthcare systems deliver better care, in-person and via telehealth, as they compensate for a growing shortage of neurologists. Current neurological digital biomarker pipelines, however, are narrowed down to a specific neurological exam component or applied for assessing specific conditions. In this paper, we propose an accessible vision-based exam and documentation solution called Digitized Neurological Examination (DNE) to expand exam biomarker recording options and clinical applications using a smartphone/tablet. Through our DNE software, healthcare providers in clinical settings and people at home are enabled to video capture an examination while performing instructed neurological tests, including finger tapping, finger to finger, forearm roll, and stand-up and walk. Our modular design of the DNE software supports integrations of additional tests. The DNE extracts from the recorded examinations the 2D/3D human-body pose and quantifies kinematic and spatio-temporal features. The features are clinically relevant and allow clinicians to document and observe the quantified movements and the changes of these metrics over time. A web server and a user interface for recordings viewing and feature visualizations are available. DNE was evaluated on a collected dataset of 21 subjects containing normal and simulated-impaired movements. The overall accuracy of DNE is demonstrated by classifying the recorded movements using various machine learning models. Our tests show an accuracy beyond 90% for upper-limb tests and 80% for the stand-up and walk tests.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge