Euijung Ryu
Extracting Social Support and Social Isolation Information from Clinical Psychiatry Notes: Comparing a Rule-based NLP System and a Large Language Model
Mar 25, 2024Abstract:Background: Social support (SS) and social isolation (SI) are social determinants of health (SDOH) associated with psychiatric outcomes. In electronic health records (EHRs), individual-level SS/SI is typically documented as narrative clinical notes rather than structured coded data. Natural language processing (NLP) algorithms can automate the otherwise labor-intensive process of data extraction. Data and Methods: Psychiatric encounter notes from Mount Sinai Health System (MSHS, n=300) and Weill Cornell Medicine (WCM, n=225) were annotated and established a gold standard corpus. A rule-based system (RBS) involving lexicons and a large language model (LLM) using FLAN-T5-XL were developed to identify mentions of SS and SI and their subcategories (e.g., social network, instrumental support, and loneliness). Results: For extracting SS/SI, the RBS obtained higher macro-averaged f-scores than the LLM at both MSHS (0.89 vs. 0.65) and WCM (0.85 vs. 0.82). For extracting subcategories, the RBS also outperformed the LLM at both MSHS (0.90 vs. 0.62) and WCM (0.82 vs. 0.81). Discussion and Conclusion: Unexpectedly, the RBS outperformed the LLMs across all metrics. Intensive review demonstrates that this finding is due to the divergent approach taken by the RBS and LLM. The RBS were designed and refined to follow the same specific rules as the gold standard annotations. Conversely, the LLM were more inclusive with categorization and conformed to common English-language understanding. Both approaches offer advantages and are made available open-source for future testing.
Neural Language Models with Distant Supervision to Identify Major Depressive Disorder from Clinical Notes
Apr 19, 2021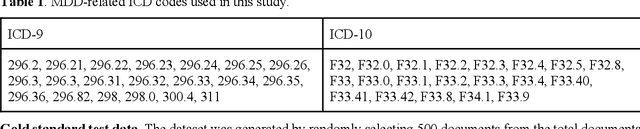
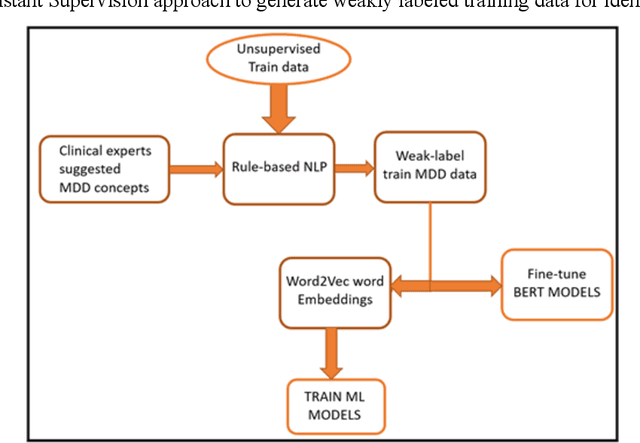
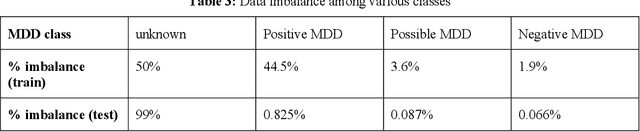
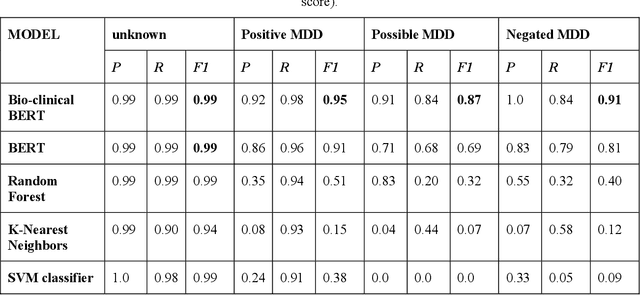
Abstract:Major depressive disorder (MDD) is a prevalent psychiatric disorder that is associated with significant healthcare burden worldwide. Phenotyping of MDD can help early diagnosis and consequently may have significant advantages in patient management. In prior research MDD phenotypes have been extracted from structured Electronic Health Records (EHR) or using Electroencephalographic (EEG) data with traditional machine learning models to predict MDD phenotypes. However, MDD phenotypic information is also documented in free-text EHR data, such as clinical notes. While clinical notes may provide more accurate phenotyping information, natural language processing (NLP) algorithms must be developed to abstract such information. Recent advancements in NLP resulted in state-of-the-art neural language models, such as Bidirectional Encoder Representations for Transformers (BERT) model, which is a transformer-based model that can be pre-trained from a corpus of unsupervised text data and then fine-tuned on specific tasks. However, such neural language models have been underutilized in clinical NLP tasks due to the lack of large training datasets. In the literature, researchers have utilized the distant supervision paradigm to train machine learning models on clinical text classification tasks to mitigate the issue of lacking annotated training data. It is still unknown whether the paradigm is effective for neural language models. In this paper, we propose to leverage the neural language models in a distant supervision paradigm to identify MDD phenotypes from clinical notes. The experimental results indicate that our proposed approach is effective in identifying MDD phenotypes and that the Bio- Clinical BERT, a specific BERT model for clinical data, achieved the best performance in comparison with conventional machine learning models.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge