Eric J. Verbeke
Fast alignment of heterogeneous images in sliced Wasserstein distance
Mar 17, 2025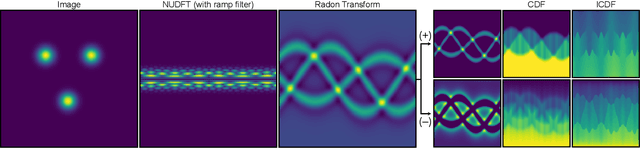

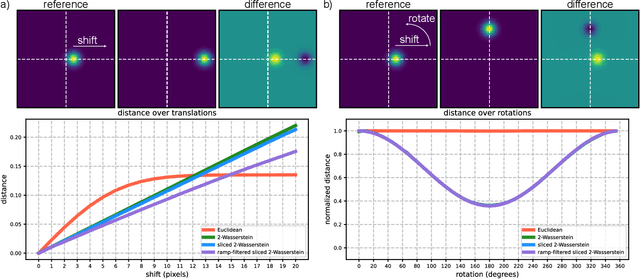
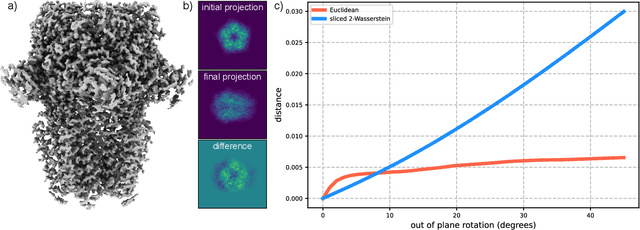
Abstract:Many applications of computer vision rely on the alignment of similar but non-identical images. We present a fast algorithm for aligning heterogeneous images based on optimal transport. Our approach combines the speed of fast Fourier methods with the robustness of sliced probability metrics and allows us to efficiently compute the alignment between two $L \times L$ images using the sliced 2-Wasserstein distance in $O(L^2 \log L)$ operations. We show that our method is robust to translations, rotations and deformations in the images.
Moment-based metrics for molecules computable from cryo-EM images
Jan 26, 2024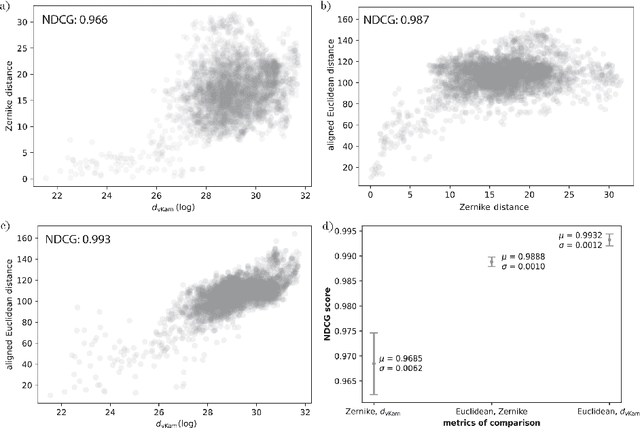
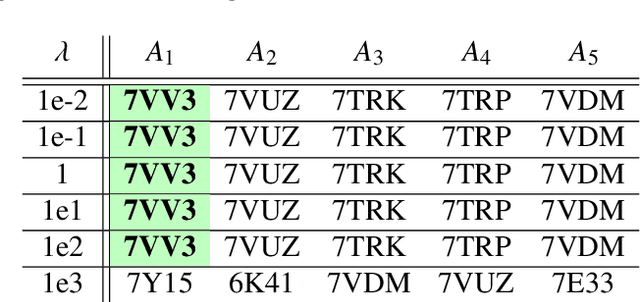
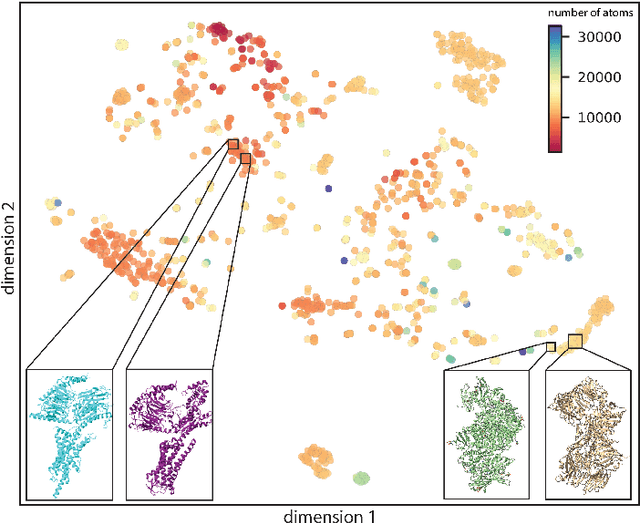
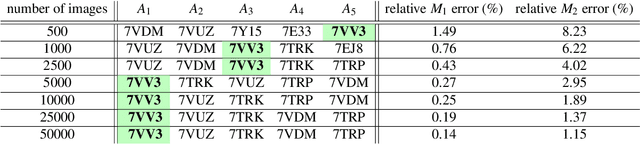
Abstract:Single particle cryogenic electron microscopy (cryo-EM) is an imaging technique capable of recovering the high-resolution 3-D structure of biological macromolecules from many noisy and randomly oriented projection images. One notable approach to 3-D reconstruction, known as Kam's method, relies on the moments of the 2-D images. Inspired by Kam's method, we introduce a rotationally invariant metric between two molecular structures, which does not require 3-D alignment. Further, we introduce a metric between a stack of projection images and a molecular structure, which is invariant to rotations and reflections and does not require performing 3-D reconstruction. Additionally, the latter metric does not assume a uniform distribution of viewing angles. We demonstrate uses of the new metrics on synthetic and experimental datasets, highlighting their ability to measure structural similarity.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge