Du Nan
CogniBench: A Legal-inspired Framework and Dataset for Assessing Cognitive Faithfulness of Large Language Models
May 28, 2025Abstract:Faithfulness hallucination are claims generated by a Large Language Model (LLM) not supported by contexts provided to the LLM. Lacking assessment standard, existing benchmarks only contain "factual statements" that rephrase source materials without marking "cognitive statements" that make inference from the given context, making the consistency evaluation and optimization of cognitive statements difficult. Inspired by how an evidence is assessed in the legislative domain, we design a rigorous framework to assess different levels of faithfulness of cognitive statements and create a benchmark dataset where we reveal insightful statistics. We design an annotation pipeline to create larger benchmarks for different LLMs automatically, and the resulting larger-scale CogniBench-L dataset can be used to train accurate cognitive hallucination detection model. We release our model and dataset at: https://github.com/FUTUREEEEEE/CogniBench
ProDOMA: improve PROtein DOMAin classification for third-generation sequencing reads using deep learning
Sep 26, 2020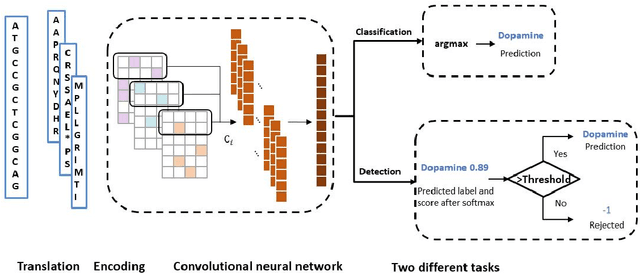
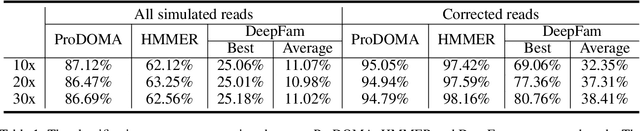
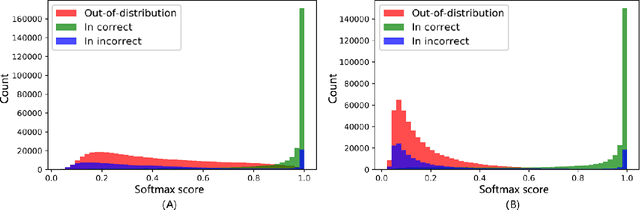

Abstract:Motivation: With the development of third-generation sequencing technologies, people are able to obtain DNA sequences with lengths from 10s to 100s of kb. These long reads allow protein domain annotation without assembly, thus can produce important insights into the biological functions of the underlying data. However, the high error rate in third-generation sequencing data raises a new challenge to established domain analysis pipelines. The state-of-the-art methods are not optimized for noisy reads and have shown unsatisfactory accuracy of domain classification in third-generation sequencing data. New computational methods are still needed to improve the performance of domain prediction in long noisy reads. Results: In this work, we introduce ProDOMA, a deep learning model that conducts domain classification for third-generation sequencing reads. It uses deep neural networks with 3-frame translation encoding to learn conserved features from partially correct translations. In addition, we formulate our problem as an open-set problem and thus our model can reject unrelated DNA reads such as those from noncoding regions. In the experiments on simulated reads of protein coding sequences and real reads from the human genome, our model outperforms HMMER and DeepFam on protein domain classification. In summary, ProDOMA is a useful end-to-end protein domain analysis tool for long noisy reads without relying on error correction. Availability: The source code and the trained model are freely available at https://github.com/strideradu/ProDOMA. Contact: yannisun@cityu.edu.hk
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge