Dedi Wang
Graph Neural Network-State Predictive Information Bottleneck (GNN-SPIB) approach for learning molecular thermodynamics and kinetics
Sep 18, 2024Abstract:Molecular dynamics simulations offer detailed insights into atomic motions but face timescale limitations. Enhanced sampling methods have addressed these challenges but even with machine learning, they often rely on pre-selected expert-based features. In this work, we present the Graph Neural Network-State Predictive Information Bottleneck (GNN-SPIB) framework, which combines graph neural networks and the State Predictive Information Bottleneck to automatically learn low-dimensional representations directly from atomic coordinates. Tested on three benchmark systems, our approach predicts essential structural, thermodynamic and kinetic information for slow processes, demonstrating robustness across diverse systems. The method shows promise for complex systems, enabling effective enhanced sampling without requiring pre-defined reaction coordinates or input features.
Introducing dynamical constraints into representation learning
Sep 02, 2022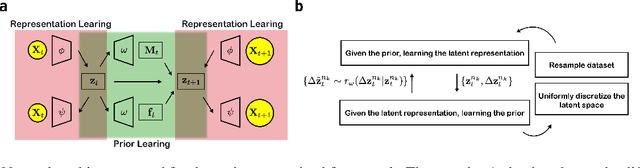
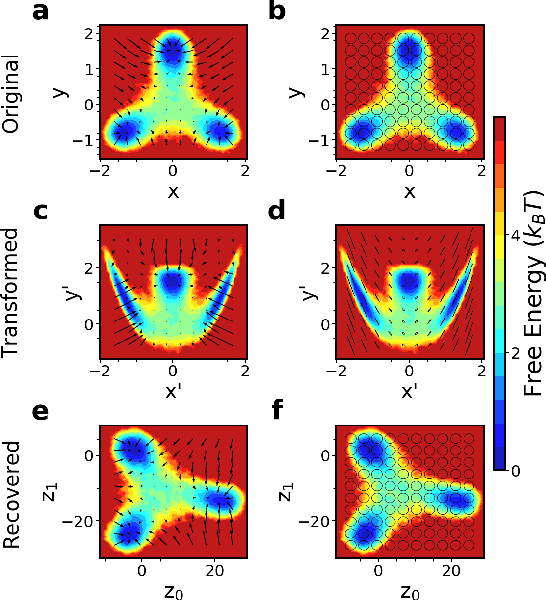

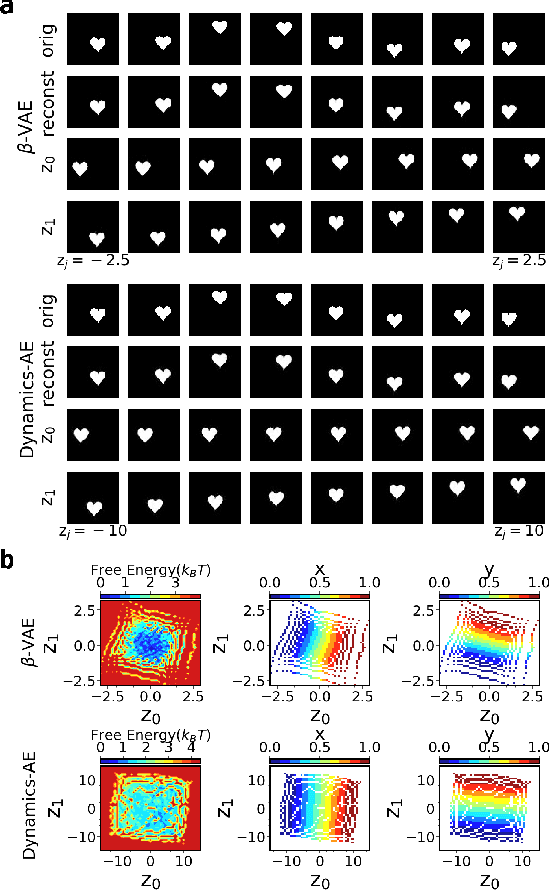
Abstract:While representation learning has been central to the rise of machine learning and artificial intelligence, a key problem remains in making the learnt representations meaningful. For this the typical approach is to regularize the learned representation through prior probability distributions. However such priors are usually unavailable or ad hoc. To deal with this, we propose a dynamics-constrained representation learning framework. Instead of using predefined probabilities, we restrict the latent representation to follow specific dynamics, which is a more natural constraint for representation learning in dynamical systems. Our belief stems from a fundamental observation in physics that though different systems can have different marginalized probability distributions, they typically obey the same dynamics, such as Newton's and Schrodinger's equations. We validate our framework for different systems including a real-world fluorescent DNA movie dataset. We show that our algorithm can uniquely identify an uncorrelated, isometric and meaningful latent representation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge