Deana Crocetti
Computerized Assessment of Motor Imitation for Distinguishing Autism in Video (CAMI-2DNet)
Jan 15, 2025Abstract:Motor imitation impairments are commonly reported in individuals with autism spectrum conditions (ASCs), suggesting that motor imitation could be used as a phenotype for addressing autism heterogeneity. Traditional methods for assessing motor imitation are subjective, labor-intensive, and require extensive human training. Modern Computerized Assessment of Motor Imitation (CAMI) methods, such as CAMI-3D for motion capture data and CAMI-2D for video data, are less subjective. However, they rely on labor-intensive data normalization and cleaning techniques, and human annotations for algorithm training. To address these challenges, we propose CAMI-2DNet, a scalable and interpretable deep learning-based approach to motor imitation assessment in video data, which eliminates the need for data normalization, cleaning and annotation. CAMI-2DNet uses an encoder-decoder architecture to map a video to a motion encoding that is disentangled from nuisance factors such as body shape and camera views. To learn a disentangled representation, we employ synthetic data generated by motion retargeting of virtual characters through the reshuffling of motion, body shape, and camera views, as well as real participant data. To automatically assess how well an individual imitates an actor, we compute a similarity score between their motion encodings, and use it to discriminate individuals with ASCs from neurotypical (NT) individuals. Our comparative analysis demonstrates that CAMI-2DNet has a strong correlation with human scores while outperforming CAMI-2D in discriminating ASC vs NT children. Moreover, CAMI-2DNet performs comparably to CAMI-3D while offering greater practicality by operating directly on video data and without the need for ad-hoc data normalization and human annotations.
A Matrix Autoencoder Framework to Align the Functional and Structural Connectivity Manifolds as Guided by Behavioral Phenotypes
May 30, 2021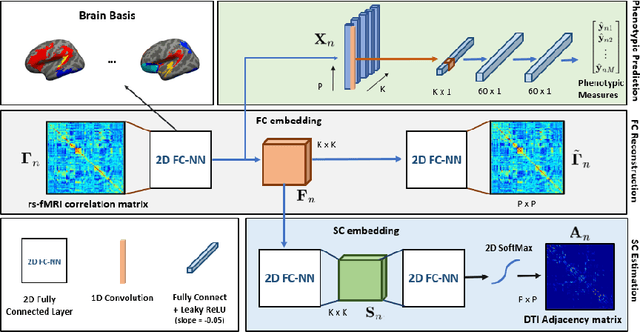
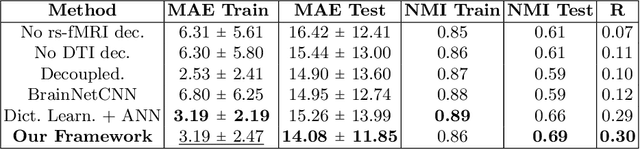
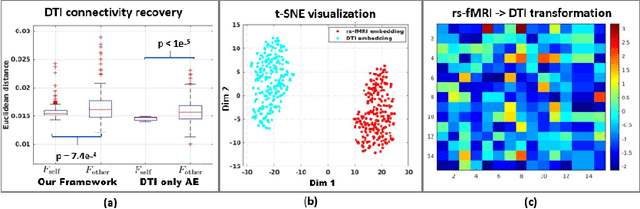
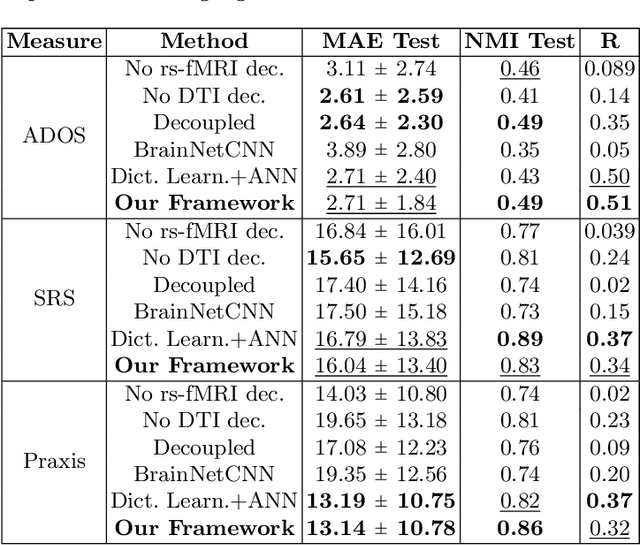
Abstract:We propose a novel matrix autoencoder to map functional connectomes from resting state fMRI (rs-fMRI) to structural connectomes from Diffusion Tensor Imaging (DTI), as guided by subject-level phenotypic measures. Our specialized autoencoder infers a low dimensional manifold embedding for the rs-fMRI correlation matrices that mimics a canonical outer-product decomposition. The embedding is simultaneously used to reconstruct DTI tractography matrices via a second manifold alignment decoder and to predict inter-subject phenotypic variability via an artificial neural network. We validate our framework on a dataset of 275 healthy individuals from the Human Connectome Project database and on a second clinical dataset consisting of 57 subjects with Autism Spectrum Disorder. We demonstrate that the model reliably recovers structural connectivity patterns across individuals, while robustly extracting predictive and interpretable brain biomarkers in a cross-validated setting. Finally, our framework outperforms several baselines at predicting behavioral phenotypes in both real-world datasets.
Deep sr-DDL: Deep Structurally Regularized Dynamic Dictionary Learning to Integrate Multimodal and Dynamic Functional Connectomics data for Multidimensional Clinical Characterizations
Aug 27, 2020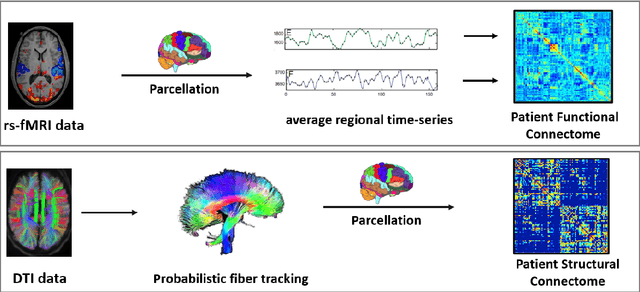
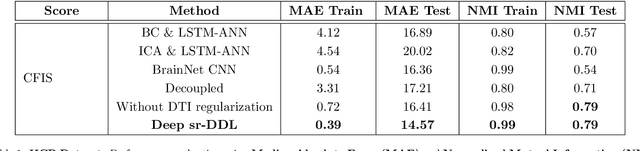
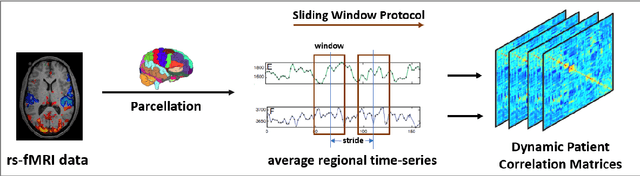
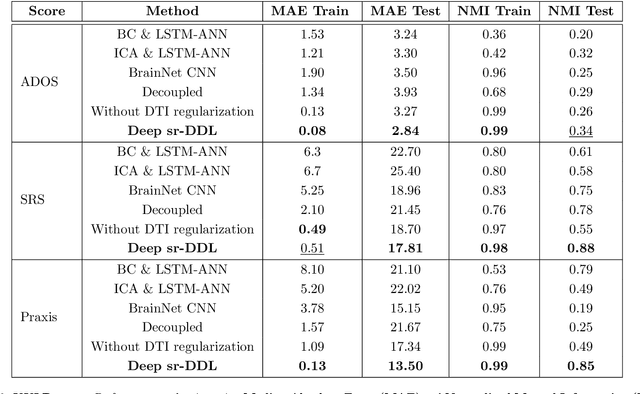
Abstract:We propose a novel integrated framework that jointly models complementary information from resting-state functional MRI (rs-fMRI) connectivity and diffusion tensor imaging (DTI) tractography to extract biomarkers of brain connectivity predictive of behavior. Our framework couples a generative model of the connectomics data with a deep network that predicts behavioral scores. The generative component is a structurally-regularized Dynamic Dictionary Learning (sr-DDL) model that decomposes the dynamic rs-fMRI correlation matrices into a collection of shared basis networks and time varying subject-specific loadings. We use the DTI tractography to regularize this matrix factorization and learn anatomically informed functional connectivity profiles. The deep component of our framework is an LSTM-ANN block, which uses the temporal evolution of the subject-specific sr-DDL loadings to predict multidimensional clinical characterizations. Our joint optimization strategy collectively estimates the basis networks, the subject-specific time-varying loadings, and the neural network weights. We validate our framework on a dataset of neurotypical individuals from the Human Connectome Project (HCP) database to map to cognition and on a separate multi-score prediction task on individuals diagnosed with Autism Spectrum Disorder (ASD) in a five-fold cross validation setting. Our hybrid model outperforms several state-of-the-art approaches at clinical outcome prediction and learns interpretable multimodal neural signatures of brain organization.
A Deep-Generative Hybrid Model to Integrate Multimodal and Dynamic Connectivity for Predicting Spectrum-Level Deficits in Autism
Jul 03, 2020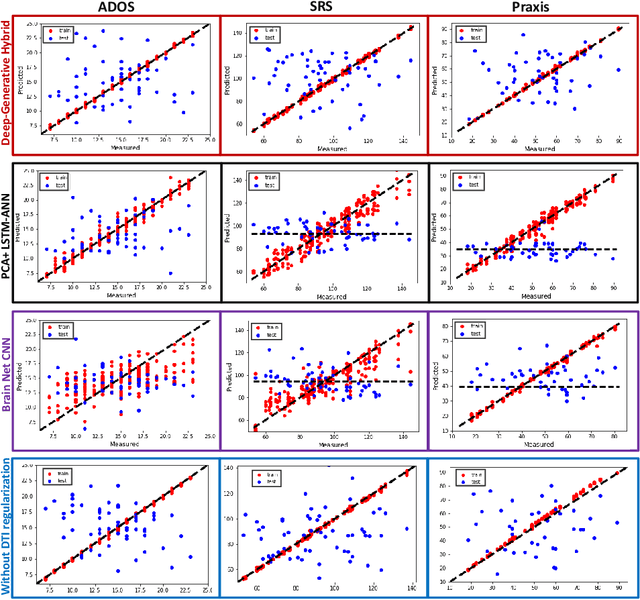
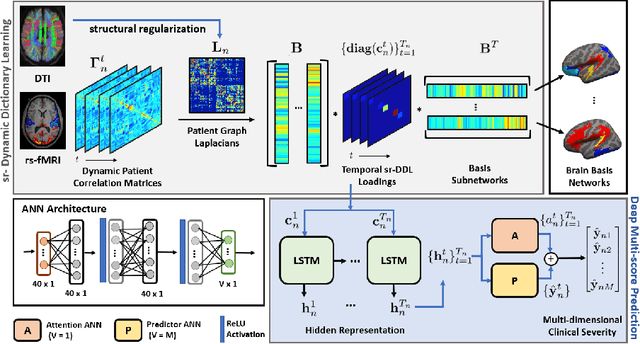
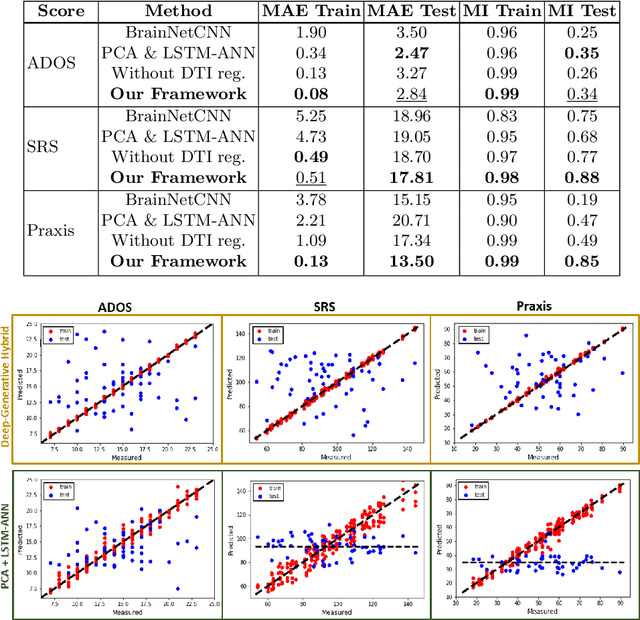
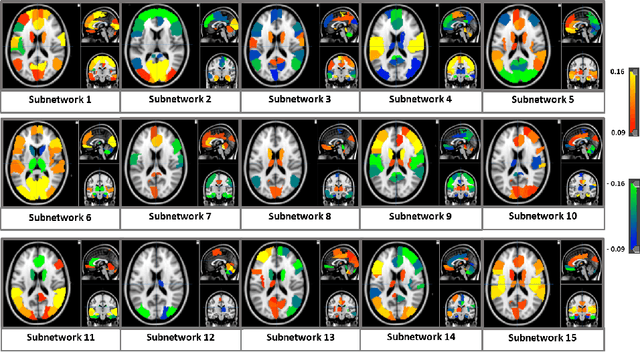
Abstract:We propose an integrated deep-generative framework, that jointly models complementary information from resting-state functional MRI (rs-fMRI) connectivity and diffusion tensor imaging (DTI) tractography to extract predictive biomarkers of a disease. The generative part of our framework is a structurally-regularized Dynamic Dictionary Learning (sr-DDL) model that decomposes the dynamic rs-fMRI correlation matrices into a collection of shared basis networks and time varying patient-specific loadings. This matrix factorization is guided by the DTI tractography matrices to learn anatomically informed connectivity profiles. The deep part of our framework is an LSTM-ANN block, which models the temporal evolution of the patient sr-DDL loadings to predict multidimensional clinical severity. Our coupled optimization procedure collectively estimates the basis networks, the patient-specific dynamic loadings, and the neural network weights. We validate our framework on a multi-score prediction task in 57 patients diagnosed with Autism Spectrum Disorder (ASD). Our hybrid model outperforms state-of-the-art baselines in a five-fold cross validated setting and extracts interpretable multimodal neural signatures of brain dysfunction in ASD.
Neuropsychiatric Disease Classification Using Functional Connectomics -- Results of the Connectomics in NeuroImaging Transfer Learning Challenge
Jun 05, 2020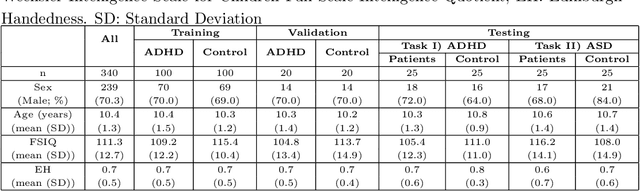
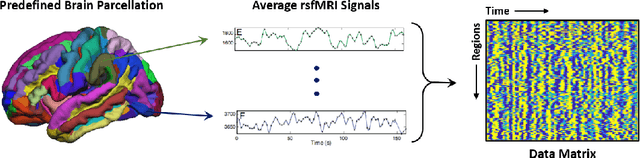
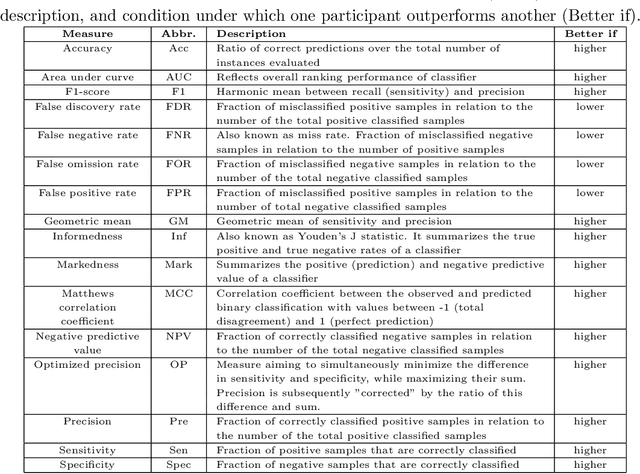
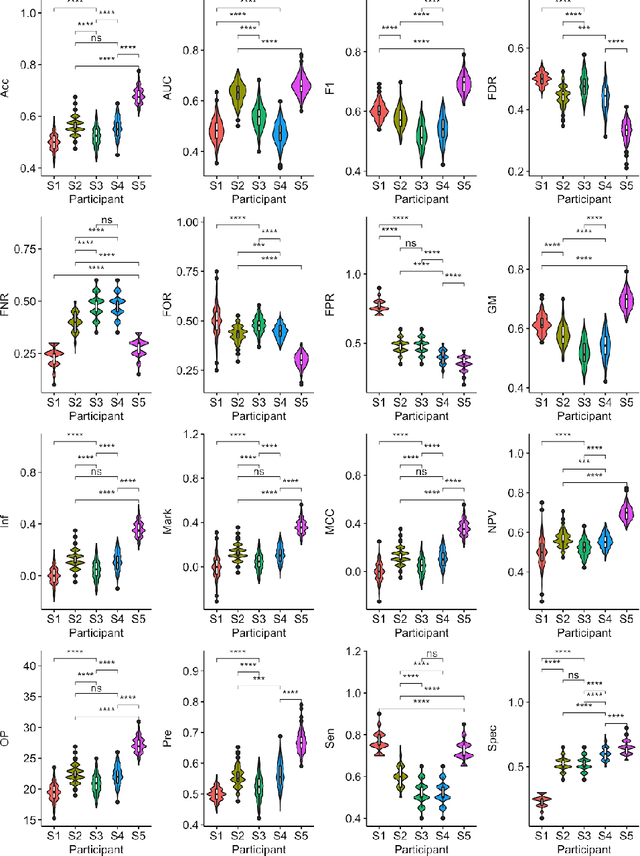
Abstract:Large, open-source consortium datasets have spurred the development of new and increasingly powerful machine learning approaches in brain connectomics. However, one key question remains: are we capturing biologically relevant and generalizable information about the brain, or are we simply overfitting to the data? To answer this, we organized a scientific challenge, the Connectomics in NeuroImaging Transfer Learning Challenge (CNI-TLC), held in conjunction with MICCAI 2019. CNI-TLC included two classification tasks: (1) diagnosis of Attention-Deficit/Hyperactivity Disorder (ADHD) within a pre-adolescent cohort; and (2) transference of the ADHD model to a related cohort of Autism Spectrum Disorder (ASD) patients with an ADHD comorbidity. In total, 240 resting-state fMRI time series averaged according to three standard parcellation atlases, along with clinical diagnosis, were released for training and validation (120 neurotypical controls and 120 ADHD). We also provided demographic information of age, sex, IQ, and handedness. A second set of 100 subjects (50 neurotypical controls, 25 ADHD, and 25 ASD with ADHD comorbidity) was used for testing. Models were submitted in a standardized format as Docker images through ChRIS, an open-source image analysis platform. Utilizing an inclusive approach, we ranked the methods based on 16 different metrics. The final rank was calculated using the rank product for each participant across all measures. Furthermore, we assessed the calibration curves of each method. Five participants submitted their model for evaluation, with one outperforming all other methods in both ADHD and ASD classification. However, further improvements are needed to reach the clinical translation of functional connectomics. We are keeping the CNI-TLC open as a publicly available resource for developing and validating new classification methodologies in the field of connectomics.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge