Christopher B. Cooper
Lung2Lung: Volumetric Style Transfer with Self-Ensembling for High-Resolution Cross-Volume Computed Tomography
Oct 06, 2022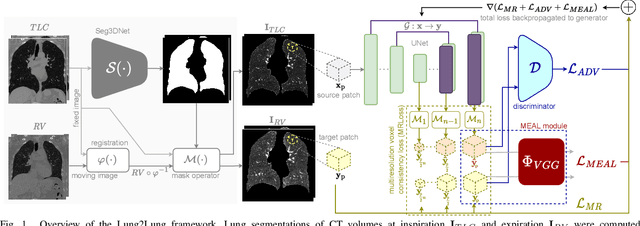
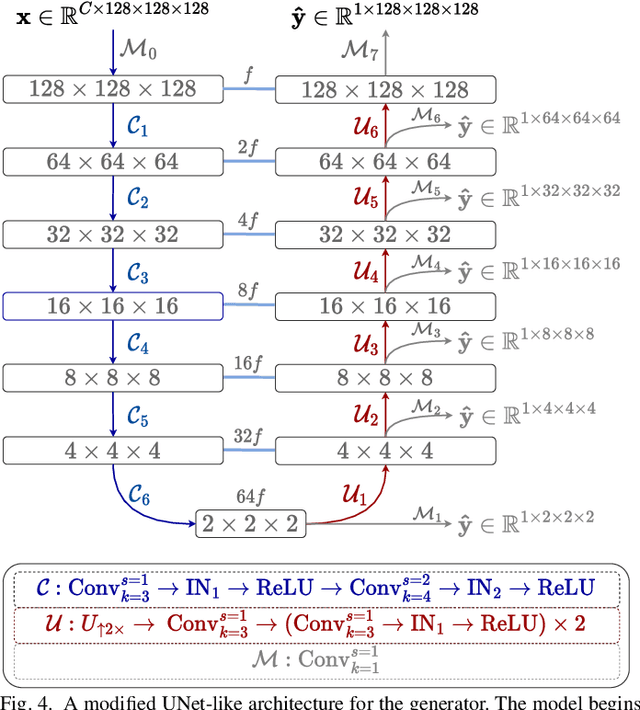
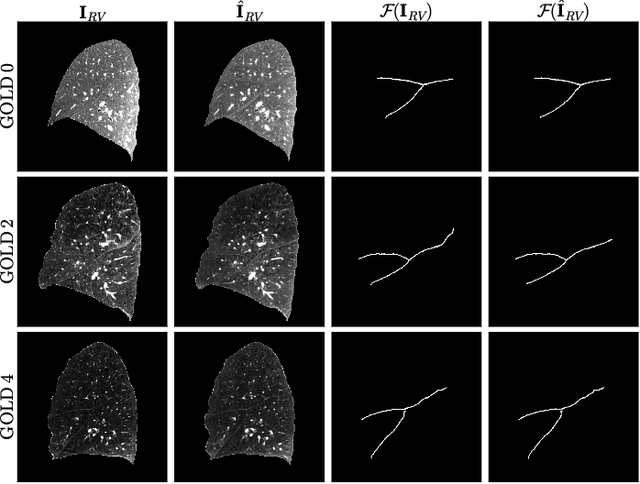
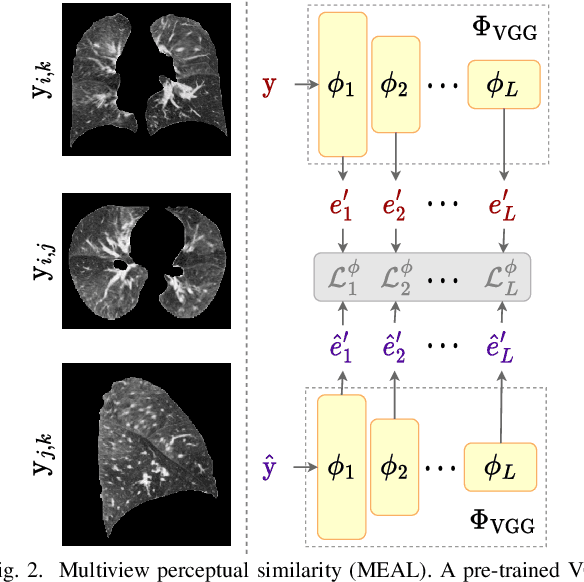

Abstract:Chest computed tomography (CT) at inspiration is often complemented by an expiratory CT for identifying peripheral airways disease in the form of air trapping. Additionally, co-registered inspiratory-expiratory volumes are used to derive several clinically relevant measures of local lung function. Acquiring CT at different volumes, however, increases radiation dosage, acquisition time, and may not be achievable due to various complications, limiting the utility of registration-based measures, To address this, we propose Lung2Lung - a style-based generative adversarial approach for translating CT images from end-inspiratory to end-expiratory volume. Lung2Lung addresses several limitations of the traditional generative models including slicewise discontinuities, limited size of generated volumes, and their inability to model neural style at a volumetric level. We introduce multiview perceptual similarity (MEAL) to capture neural styles in 3D. To incorporate global information into the training process and refine the output of our model, we also propose self-ensembling (SE). Lung2Lung, with MEAL and SE, is able to generate large 3D volumes of size 320 x 320 x 320 that are validated using a diverse cohort of 1500 subjects with varying disease severity. The model shows superior performance against several state-of-the-art 2D and 3D generative models with a peak-signal-to-noise ratio of 24.53 dB and structural similarity of 0.904. Clinical validation shows that the synthetic volumes can be used to reliably extract several clinical endpoints of chronic obstructive pulmonary disease.
Single volume lung biomechanics from chest computed tomography using a mode preserving generative adversarial network
Oct 15, 2021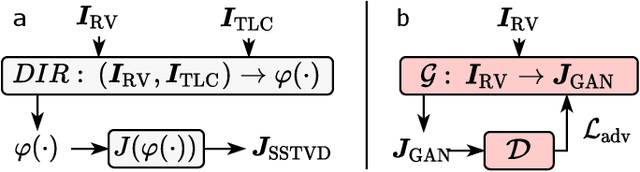
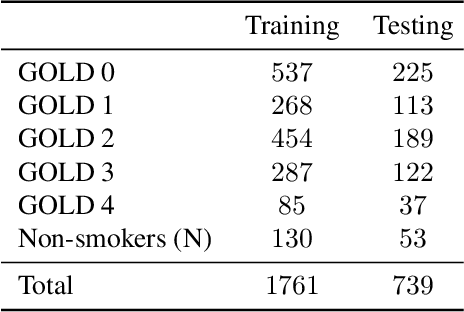
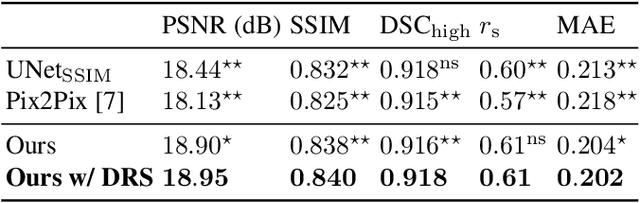
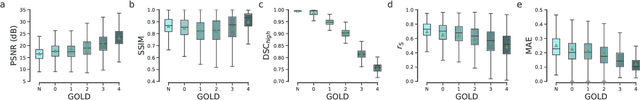
Abstract:Local tissue expansion of the lungs is typically derived by registering computed tomography (CT) scans acquired at multiple lung volumes. However, acquiring multiple scans incurs increased radiation dose, time, and cost, and may not be possible in many cases, thus restricting the applicability of registration-based biomechanics. We propose a generative adversarial learning approach for estimating local tissue expansion directly from a single CT scan. The proposed framework was trained and evaluated on 2500 subjects from the SPIROMICS cohort. Once trained, the framework can be used as a registration-free method for predicting local tissue expansion. We evaluated model performance across varying degrees of disease severity and compared its performance with two image-to-image translation frameworks - UNet and Pix2Pix. Our model achieved an overall PSNR of 18.95 decibels, SSIM of 0.840, and Spearman's correlation of 0.61 at a high spatial resolution of 1 mm3.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge