Christof Bertram
Quantifying the Scanner-Induced Domain Gap in Mitosis Detection
Mar 30, 2021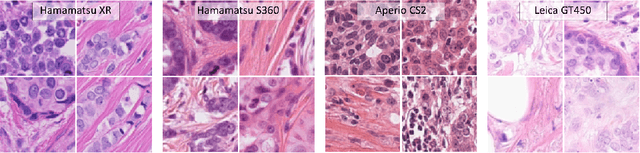
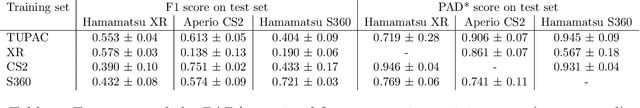

Abstract:Automated detection of mitotic figures in histopathology images has seen vast improvements, thanks to modern deep learning-based pipelines. Application of these methods, however, is in practice limited by strong variability of images between labs. This results in a domain shift of the images, which causes a performance drop of the models. Hypothesizing that the scanner device plays a decisive role in this effect, we evaluated the susceptibility of a standard mitosis detection approach to the domain shift introduced by using a different whole slide scanner. Our work is based on the MICCAI-MIDOG challenge 2021 data set, which includes 200 tumor cases of human breast cancer and four scanners. Our work indicates that the domain shift induced not by biochemical variability but purely by the choice of acquisition device is underestimated so far. Models trained on images of the same scanner yielded an average F1 score of 0.683, while models trained on a single other scanner only yielded an average F1 score of 0.325. Training on another multi-domain mitosis dataset led to mean F1 scores of 0.52. We found this not to be reflected by domain-shifts measured as proxy A distance-derived metric.
SlideRunner - A Tool for Massive Cell Annotations in Whole Slide Images
Feb 07, 2018
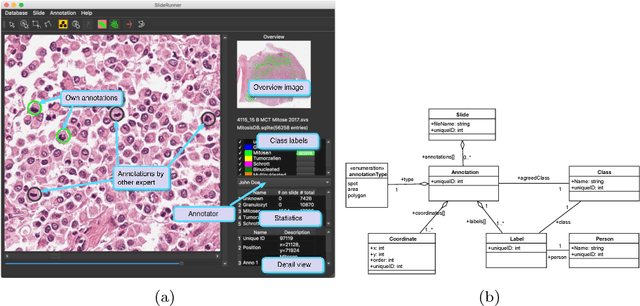
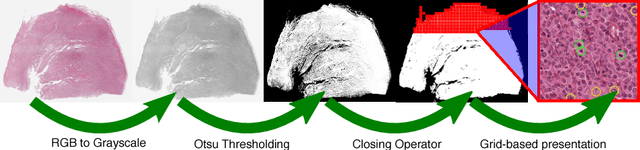
Abstract:Large-scale image data such as digital whole-slide histology images pose a challenging task at annotation software solutions. Today, a number of good solutions with varying scopes exist. For cell annotation, however, we find that many do not match the prerequisites for fast annotations. Especially in the field of mitosis detection, it is assumed that detection accuracy could significantly benefit from larger annotation databases that are currently however very troublesome to produce. Further, multiple independent (blind) expert labels are a big asset for such databases, yet there is currently no tool for this kind of annotation available. To ease this tedious process of expert annotation and grading, we introduce SlideRunner, an open source annotation and visualization tool for digital histopathology, developed in close cooperation with two pathologists. SlideRunner is capable of setting annotations like object centers (for e.g. cells) as well as object boundaries (e.g. for tumor outlines). It provides single-click annotations as well as a blind mode for multi-annotations, where the expert is directly shown the microscopy image containing the cells that he has not yet rated.
* 6 pages, submitted to Bildverarbeitung in der Medizin 2018
A Guided Spatial Transformer Network for Histology Cell Differentiation
Jul 26, 2017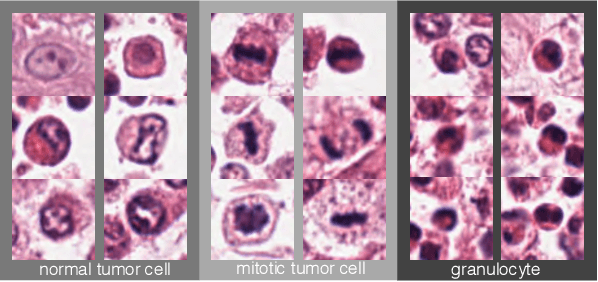
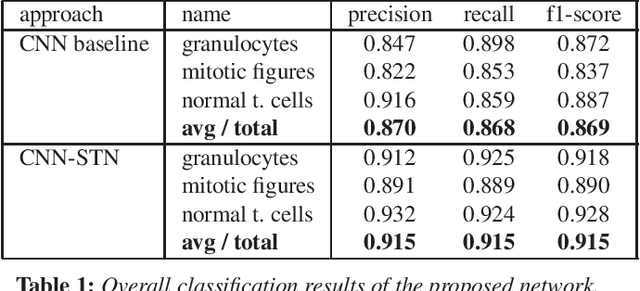
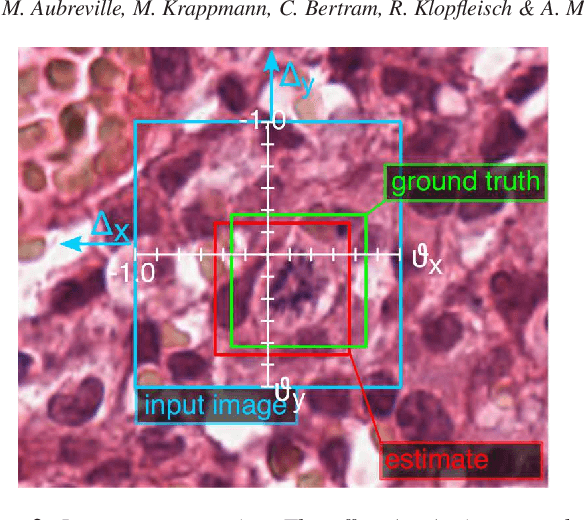
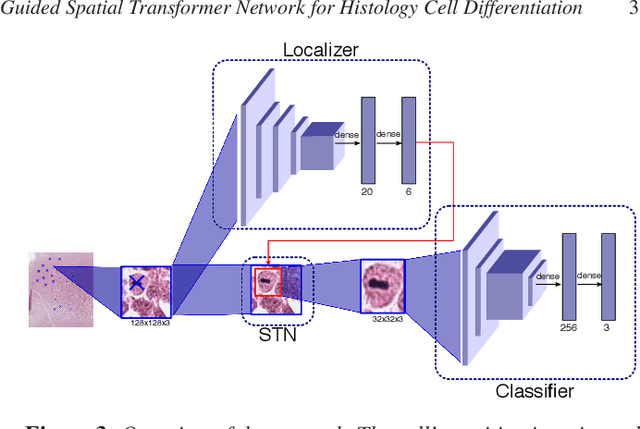
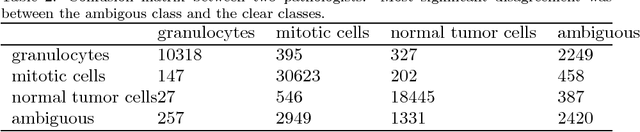
Abstract:Identification and counting of cells and mitotic figures is a standard task in diagnostic histopathology. Due to the large overall cell count on histological slides and the potential sparse prevalence of some relevant cell types or mitotic figures, retrieving annotation data for sufficient statistics is a tedious task and prone to a significant error in assessment. Automatic classification and segmentation is a classic task in digital pathology, yet it is not solved to a sufficient degree. We present a novel approach for cell and mitotic figure classification, based on a deep convolutional network with an incorporated Spatial Transformer Network. The network was trained on a novel data set with ten thousand mitotic figures, about ten times more than previous data sets. The algorithm is able to derive the cell class (mitotic tumor cells, non-mitotic tumor cells and granulocytes) and their position within an image. The mean accuracy of the algorithm in a five-fold cross-validation is 91.45%. In our view, the approach is a promising step into the direction of a more objective and accurate, semi-automatized mitosis counting supporting the pathologist.
* 5 pages, 4 figures, EG VCBM 2017 contribution
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge