Bing Yao
Geometry-aware Active Learning of Spatiotemporal Dynamic Systems
May 01, 2025
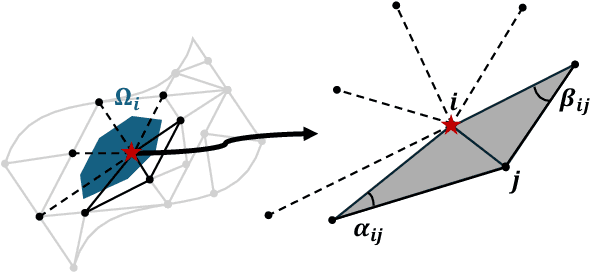
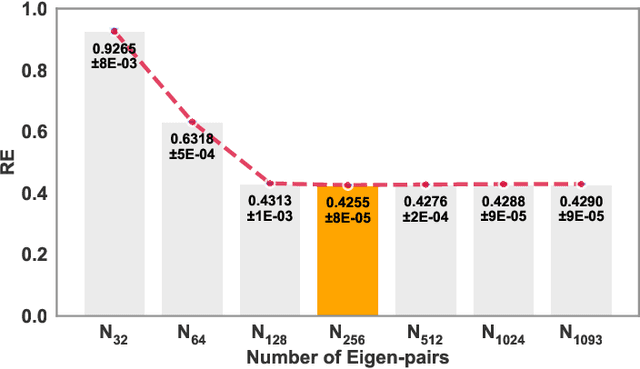
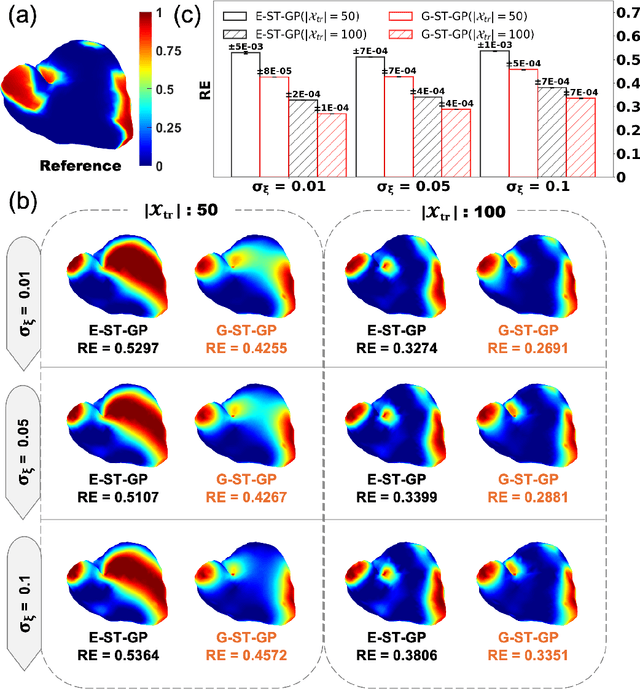
Abstract:Rapid developments in advanced sensing and imaging have significantly enhanced information visibility, opening opportunities for predictive modeling of complex dynamic systems. However, sensing signals acquired from such complex systems are often distributed across 3D geometries and rapidly evolving over time, posing significant challenges in spatiotemporal predictive modeling. This paper proposes a geometry-aware active learning framework for modeling spatiotemporal dynamic systems. Specifically, we propose a geometry-aware spatiotemporal Gaussian Process (G-ST-GP) to effectively integrate the temporal correlations and geometric manifold features for reliable prediction of high-dimensional dynamic behaviors. In addition, we develop an adaptive active learning strategy to strategically identify informative spatial locations for data collection and further maximize the prediction accuracy. This strategy achieves the adaptive trade-off between the prediction uncertainty in the G-ST-GP model and the space-filling design guided by the geodesic distance across the 3D geometry. We implement the proposed framework to model the spatiotemporal electrodynamics in a 3D heart geometry. Numerical experiments show that our framework outperforms traditional methods lacking the mechanism of geometric information incorporation or effective data collection.
SpeHeatal: A Cluster-Enhanced Segmentation Method for Sperm Morphology Analysis
Feb 18, 2025Abstract:The accurate assessment of sperm morphology is crucial in andrological diagnostics, where the segmentation of sperm images presents significant challenges. Existing approaches frequently rely on large annotated datasets and often struggle with the segmentation of overlapping sperm and the presence of dye impurities. To address these challenges, this paper first analyzes the issue of overlapping sperm tails from a geometric perspective and introduces a novel clustering algorithm, Con2Dis, which effectively segments overlapping tails by considering three essential factors: CONnectivity, CONformity, and DIStance. Building on this foundation, we propose an unsupervised method, SpeHeatal, designed for the comprehensive segmentation of the SPErm HEAd and TAiL. SpeHeatal employs the Segment Anything Model(SAM) to generate masks for sperm heads while filtering out dye impurities, utilizes Con2Dis to segment tails, and then applies a tailored mask splicing technique to produce complete sperm masks. Experimental results underscore the superior performance of SpeHeatal, particularly in handling images with overlapping sperm.
Physics-augmented Deep Learning with Adversarial Domain Adaptation: Applications to STM Image Denoising
Sep 08, 2024Abstract:Image denoising is a critical task in various scientific fields such as medical imaging and material characterization, where the accurate recovery of underlying structures from noisy data is essential. Although supervised denoising techniques have achieved significant advancements, they typically require large datasets of paired clean-noisy images for training. Unsupervised methods, while not reliant on paired data, typically necessitate a set of unpaired clean images for training, which are not always accessible. In this paper, we propose a physics-augmented deep learning with adversarial domain adaption (PDA-Net) framework for unsupervised image denoising, with applications to denoise real-world scanning tunneling microscopy (STM) images. Our PDA-Net leverages the underlying physics to simulate and envision the ground truth for denoised STM images. Additionally, built upon Generative Adversarial Networks (GANs), we incorporate a cycle-consistency module and a domain adversarial module into our PDA-Net to address the challenge of lacking paired training data and achieve information transfer between the simulated and real experimental domains. Finally, we propose to implement feature alignment and weight-sharing techniques to fully exploit the similarity between simulated and real experimental images, thereby enhancing the denoising performance in both the simulation and experimental domains. Experimental results demonstrate that the proposed PDA-Net successfully enhances the quality of STM images, offering promising applications to enhance scientific discovery and accelerate experimental quantum material research.
MUSE-Net: Missingness-aware mUlti-branching Self-attention Encoder for Irregular Longitudinal Electronic Health Records
Jun 30, 2024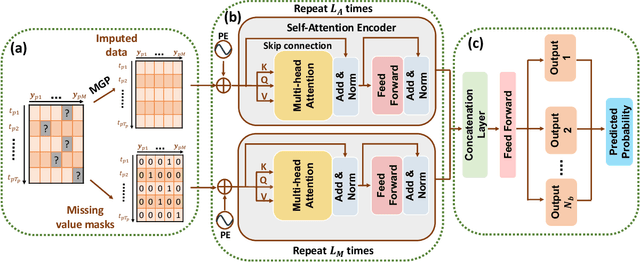
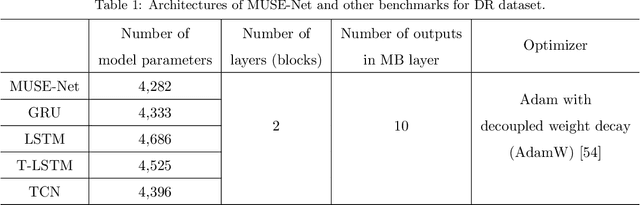

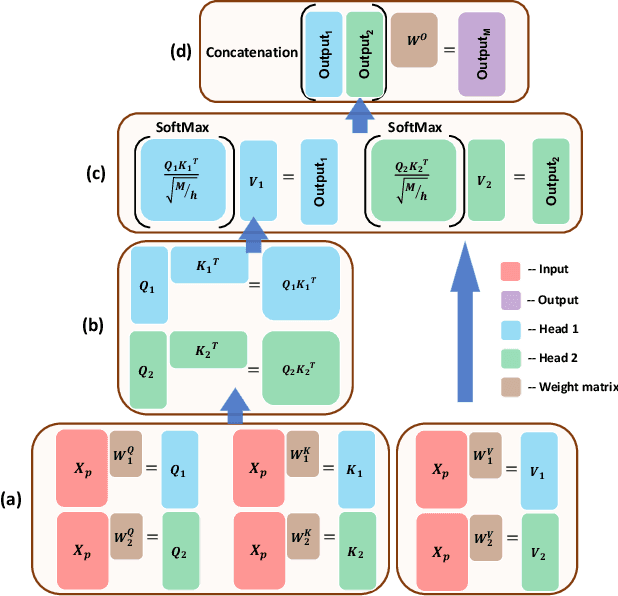
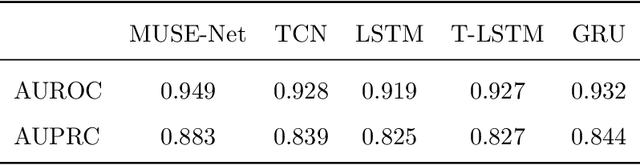
Abstract:The era of big data has made vast amounts of clinical data readily available, particularly in the form of electronic health records (EHRs), which provides unprecedented opportunities for developing data-driven diagnostic tools to enhance clinical decision making. However, the application of EHRs in data-driven modeling faces challenges such as irregularly spaced multi-variate time series, issues of incompleteness, and data imbalance. Realizing the full data potential of EHRs hinges on the development of advanced analytical models. In this paper, we propose a novel Missingness-aware mUlti-branching Self-attention Encoder (MUSE-Net) to cope with the challenges in modeling longitudinal EHRs for data-driven disease prediction. The MUSE-Net leverages a multi-task Gaussian process (MGP) with missing value masks for data imputation, a multi-branching architecture to address the data imbalance problem, and a time-aware self-attention encoder to account for the irregularly spaced time interval in longitudinal EHRs. We evaluate the proposed MUSE-Net using both synthetic and real-world datasets. Experimental results show that our MUSE-Net outperforms existing methods that are widely used to investigate longitudinal signals.
The Effect of Different Optimization Strategies to Physics-Constrained Deep Learning for Soil Moisture Estimation
Mar 13, 2024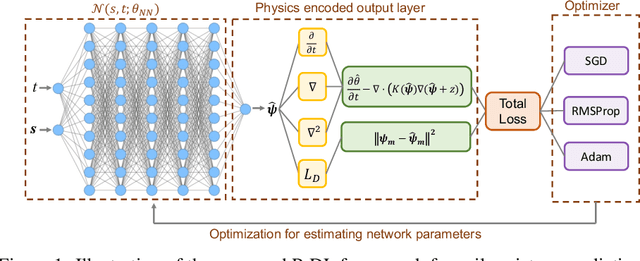
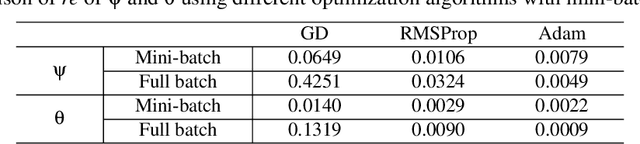
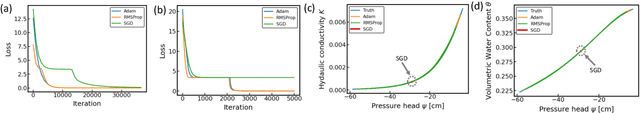
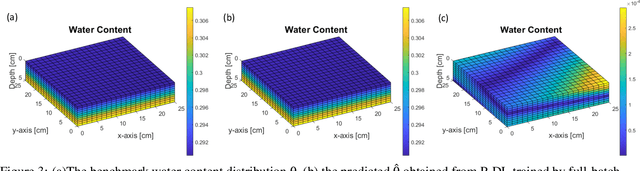
Abstract:Soil moisture is a key hydrological parameter that has significant importance to human society and the environment. Accurate modeling and monitoring of soil moisture in crop fields, especially in the root zone (top 100 cm of soil), is essential for improving agricultural production and crop yield with the help of precision irrigation and farming tools. Realizing the full sensor data potential depends greatly on advanced analytical and predictive domain-aware models. In this work, we propose a physics-constrained deep learning (P-DL) framework to integrate physics-based principles on water transport and water sensing signals for effective reconstruction of the soil moisture dynamics. We adopt three different optimizers, namely Adam, RMSprop, and GD, to minimize the loss function of P-DL during the training process. In the illustrative case study, we demonstrate the empirical convergence of Adam optimizers outperforms the other optimization methods in both mini-batch and full-batch training.
Physics-constrained Active Learning for Soil Moisture Estimation and Optimal Sensor Placement
Mar 12, 2024Abstract:Soil moisture is a crucial hydrological state variable that has significant importance to the global environment and agriculture. Precise monitoring of soil moisture in crop fields is critical to reducing agricultural drought and improving crop yield. In-situ soil moisture sensors, which are buried at pre-determined depths and distributed across the field, are promising solutions for monitoring soil moisture. However, high-density sensor deployment is neither economically feasible nor practical. Thus, to achieve a higher spatial resolution of soil moisture dynamics using a limited number of sensors, we integrate a physics-based agro-hydrological model based on Richards' equation in a physics-constrained deep learning framework to accurately predict soil moisture dynamics in the soil's root zone. This approach ensures that soil moisture estimates align well with sensor observations while obeying physical laws at the same time. Furthermore, to strategically identify the locations for sensor placement, we introduce a novel active learning framework that combines space-filling design and physics residual-based sampling to maximize data acquisition potential with limited sensors. Our numerical results demonstrate that integrating Physics-constrained Deep Learning (P-DL) with an active learning strategy within a unified framework--named the Physics-constrained Active Learning (P-DAL) framework--significantly improves the predictive accuracy and effectiveness of field-scale soil moisture monitoring using in-situ sensors.
Automated Identication of Atrial Fibrillation from Single-lead ECGs Using Multi-branching ResNet
Jun 26, 2023



Abstract:Atrial fibrillation (AF) is the most common cardiac arrhythmia, which is clinically identified with irregular and rapid heartbeat rhythm. AF puts a patient at risk of forming blood clots, which can eventually lead to heart failure, stroke, or even sudden death. It is of critical importance to develop an advanced analytical model that can effectively interpret the electrocardiography (ECG) signals and provide decision support for accurate AF diagnostics. In this paper, we propose an innovative deep-learning method for automated AF identification from single-lead ECGs. We first engage the continuous wavelet transform (CWT) to extract time-frequency features from ECG signals. Then, we develop a convolutional neural network (CNN) structure that incorporates ResNet for effective network training and multi-branching architectures for addressing the imbalanced data issue to process the 2D time-frequency features for AF classification. We evaluate the proposed methodology using two real-world ECG databases. The experimental results show a superior performance of our method compared with traditional deep learning models.
Hierarchical Deep Learning with Generative Adversarial Network for Automatic Cardiac Diagnosis from ECG Signals
Oct 19, 2022
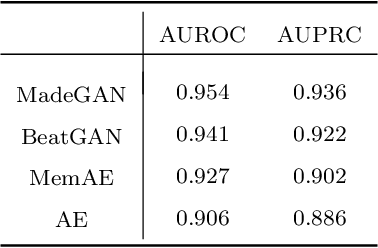

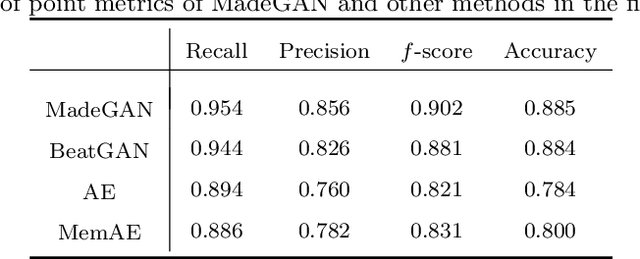
Abstract:Cardiac disease is the leading cause of death in the US. Accurate heart disease detection is of critical importance for timely medical treatment to save patients' lives. Routine use of electrocardiogram (ECG) is the most common method for physicians to assess the electrical activities of the heart and detect possible abnormal cardiac conditions. Fully utilizing the ECG data for reliable heart disease detection depends on developing effective analytical models. In this paper, we propose a two-level hierarchical deep learning framework with Generative Adversarial Network (GAN) for automatic diagnosis of ECG signals. The first-level model is composed of a Memory-Augmented Deep auto-Encoder with GAN (MadeGAN), which aims to differentiate abnormal signals from normal ECGs for anomaly detection. The second-level learning aims at robust multi-class classification for different arrhythmias identification, which is achieved by integrating the transfer learning technique to transfer knowledge from the first-level learning with the multi-branching architecture to handle the data-lacking and imbalanced data issue. We evaluate the performance of the proposed framework using real-world medical data from the MIT-BIH arrhythmia database. Experimental results show that our proposed model outperforms existing methods that are commonly used in current practice.
Physics-constrained Deep Learning for Robust Inverse ECG Modeling
Jul 26, 2021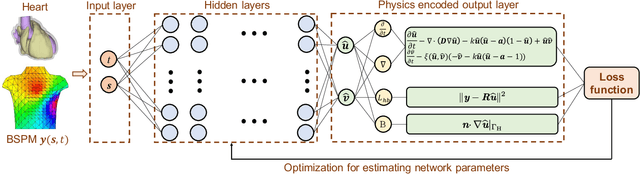
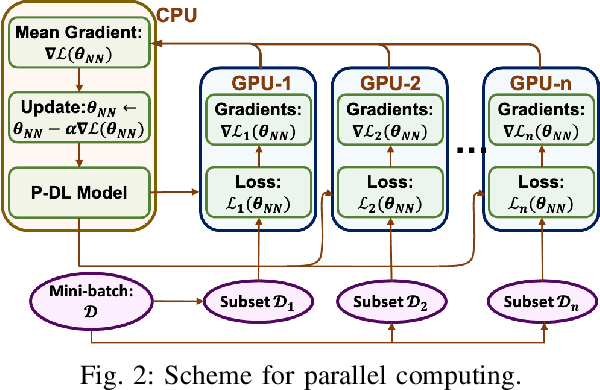
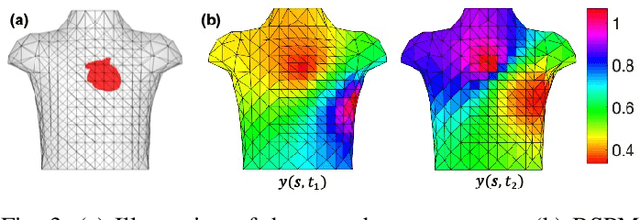
Abstract:The rapid developments in advanced sensing and imaging bring about a data-rich environment, facilitating the effective modeling, monitoring, and control of complex systems. For example, the body-sensor network captures multi-channel information pertinent to the electrical activity of the heart (i.e., electrocardiograms (ECG)), which enables medical scientists to monitor and detect abnormal cardiac conditions. However, the high-dimensional sensing data are generally complexly structured and realizing the full data potential depends to a great extent on advanced analytical and predictive methods. This paper presents a physics-constrained deep learning (P-DL) framework for high-dimensional inverse ECG modeling. This method integrates the physical laws of the complex system with the advanced deep learning infrastructure for effective prediction of the system dynamics. The proposed P-DL approach is implemented to solve the inverse ECG model and predict the time-varying distribution of electric potentials in the heart from the ECG data measured by the body-surface sensor network. Experimental results show that the proposed P-DL method significantly outperforms existing methods that are commonly used in current practice.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge