Arian Rokkum Jamasb
Structure-aware generation of drug-like molecules
Nov 07, 2021
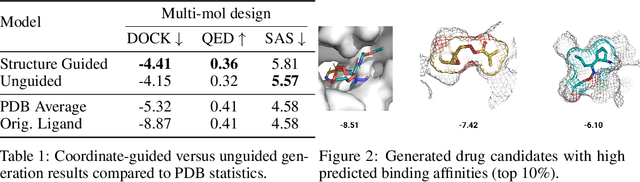
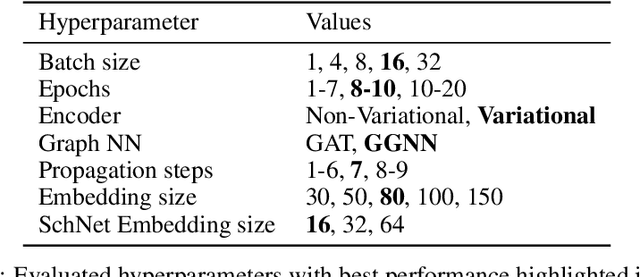
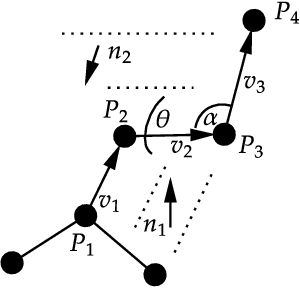
Abstract:Structure-based drug design involves finding ligand molecules that exhibit structural and chemical complementarity to protein pockets. Deep generative methods have shown promise in proposing novel molecules from scratch (de-novo design), avoiding exhaustive virtual screening of chemical space. Most generative de-novo models fail to incorporate detailed ligand-protein interactions and 3D pocket structures. We propose a novel supervised model that generates molecular graphs jointly with 3D pose in a discretised molecular space. Molecules are built atom-by-atom inside pockets, guided by structural information from crystallographic data. We evaluate our model using a docking benchmark and find that guided generation improves predicted binding affinities by 8% and drug-likeness scores by 10% over the baseline. Furthermore, our model proposes molecules with binding scores exceeding some known ligands, which could be useful in future wet-lab studies.
On Graph Neural Network Ensembles for Large-Scale Molecular Property Prediction
Jun 29, 2021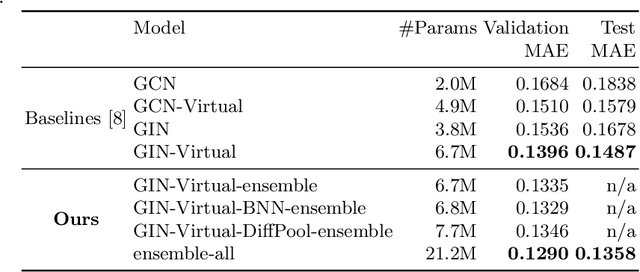
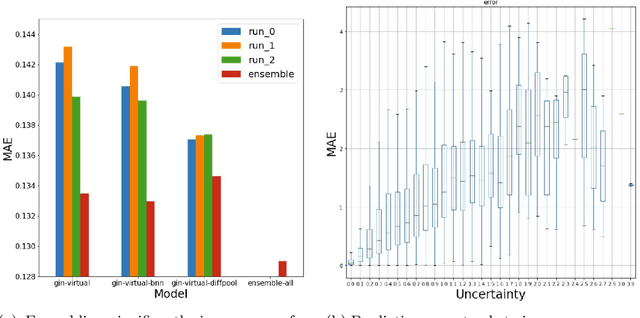
Abstract:In order to advance large-scale graph machine learning, the Open Graph Benchmark Large Scale Challenge (OGB-LSC) was proposed at the KDD Cup 2021. The PCQM4M-LSC dataset defines a molecular HOMO-LUMO property prediction task on about 3.8M graphs. In this short paper, we show our current work-in-progress solution which builds an ensemble of three graph neural networks models based on GIN, Bayesian Neural Networks and DiffPool. Our approach outperforms the provided baseline by 7.6%. Moreover, using uncertainty in our ensemble's prediction, we can identify molecules whose HOMO-LUMO gaps are harder to predict (with Pearson's correlation of 0.5181). We anticipate that this will facilitate active learning.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge