Alexander Kurz
On the calibration of neural networks for histological slide-level classification
Dec 15, 2023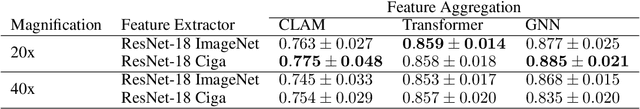
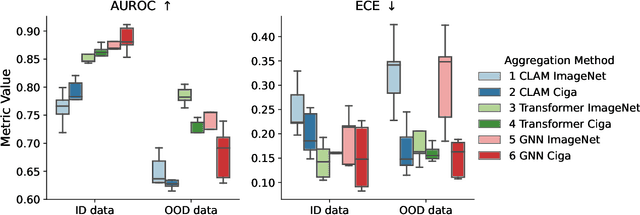
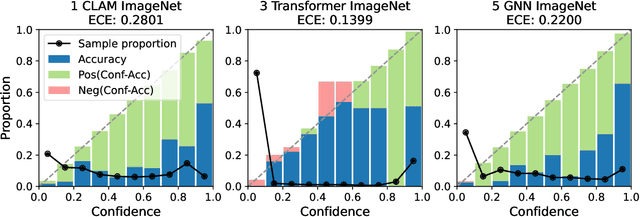
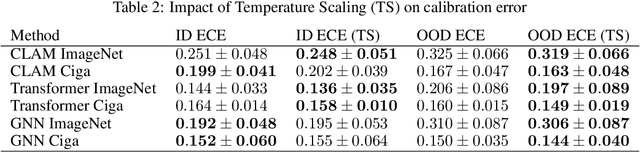
Abstract:Deep Neural Networks have shown promising classification performance when predicting certain biomarkers from Whole Slide Images in digital pathology. However, the calibration of the networks' output probabilities is often not evaluated. Communicating uncertainty by providing reliable confidence scores is of high relevance in the medical context. In this work, we compare three neural network architectures that combine feature representations on patch-level to a slide-level prediction with respect to their classification performance and evaluate their calibration. As slide-level classification task, we choose the prediction of Microsatellite Instability from Colorectal Cancer tissue sections. We observe that Transformers lead to good results in terms of classification performance and calibration. When evaluating the classification performance on a separate dataset, we observe that Transformers generalize best. The investigation of reliability diagrams provides additional insights to the Expected Calibration Error metric and we observe that especially Transformers push the output probabilities to extreme values, which results in overconfident predictions.
Benchmarking common uncertainty estimation methods with histopathological images under domain shift and label noise
Jan 03, 2023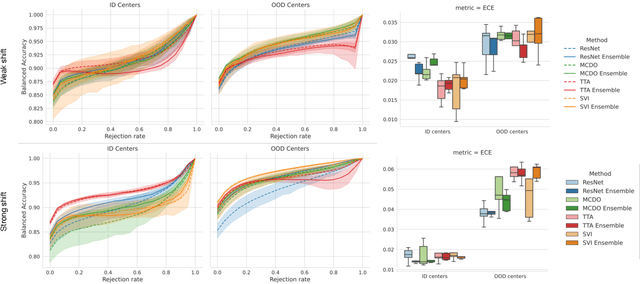
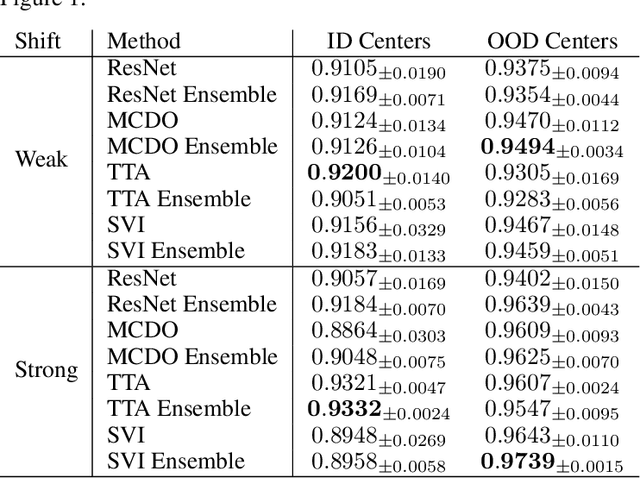
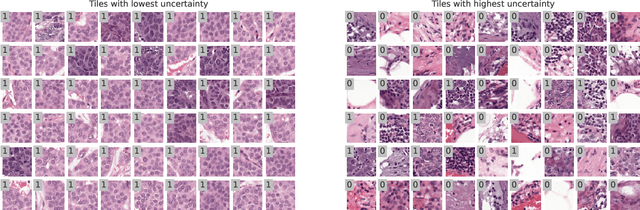
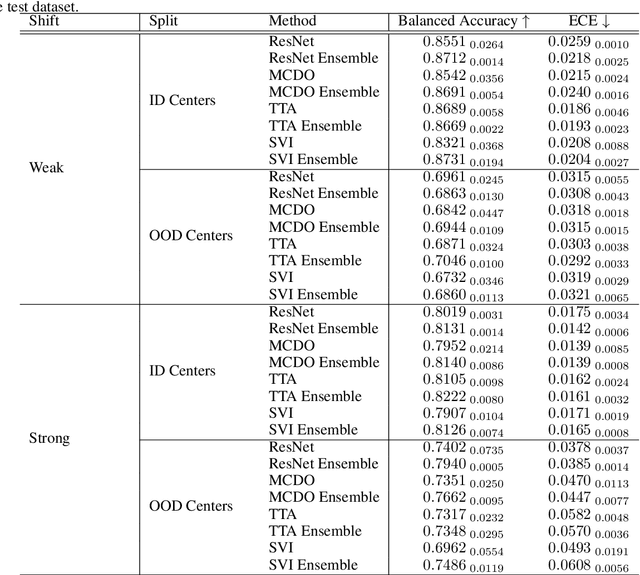
Abstract:In the past years, deep learning has seen an increase of usage in the domain of histopathological applications. However, while these approaches have shown great potential, in high-risk environments deep learning models need to be able to judge their own uncertainty and be able to reject inputs when there is a significant chance of misclassification. In this work, we conduct a rigorous evaluation of the most commonly used uncertainty and robustness methods for the classification of Whole-Slide-Images under domain shift using the H\&E stained Camelyon17 breast cancer dataset. Although it is known that histopathological data can be subject to strong domain shift and label noise, to our knowledge this is the first work that compares the most common methods for uncertainty estimation under these aspects. In our experiments, we compare Stochastic Variational Inference, Monte-Carlo Dropout, Deep Ensembles, Test-Time Data Augmentation as well as combinations thereof. We observe that ensembles of methods generally lead to higher accuracies and better calibration and that Test-Time Data Augmentation can be a promising alternative when choosing an appropriate set of augmentations. Across methods, a rejection of the most uncertain tiles leads to a significant increase in classification accuracy on both in-distribution as well as out-of-distribution data. Furthermore, we conduct experiments comparing these methods under varying conditions of label noise. We observe that the border regions of the Camelyon17 dataset are subject to label noise and evaluate the robustness of the included methods against different noise levels. Lastly, we publish our code framework to facilitate further research on uncertainty estimation on histopathological data.
Semantic Image Alignment for Vehicle Localization
Oct 08, 2021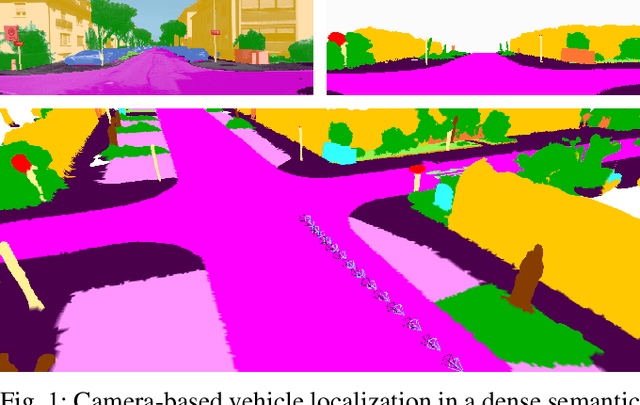
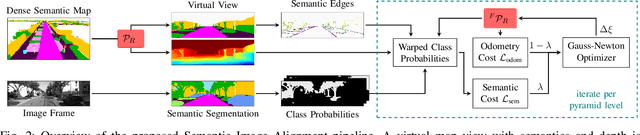
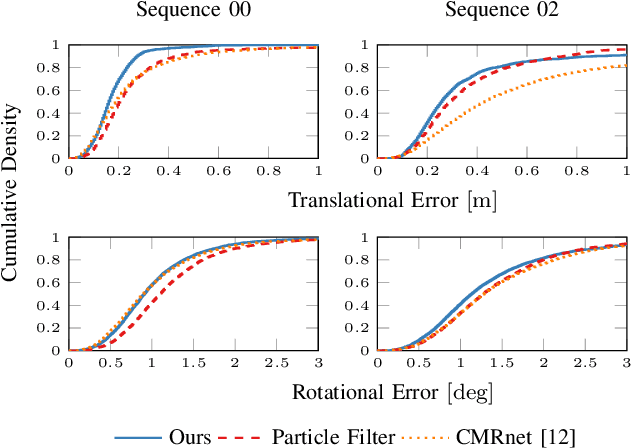
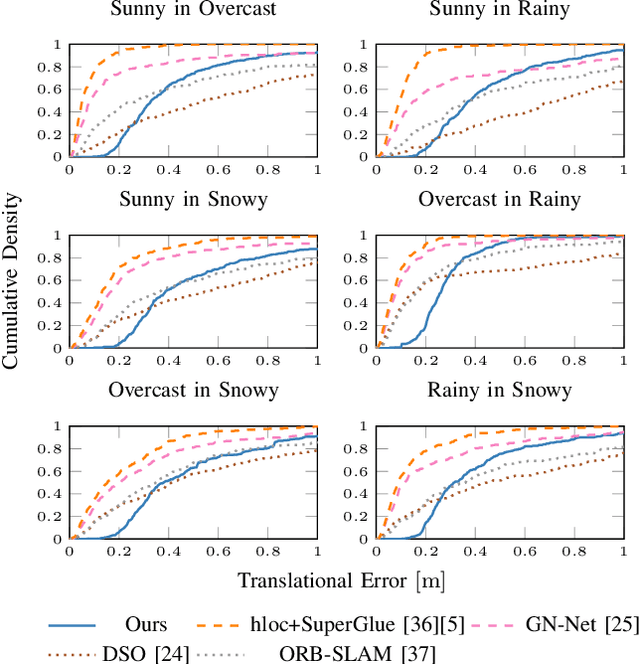
Abstract:Accurate and reliable localization is a fundamental requirement for autonomous vehicles to use map information in higher-level tasks such as navigation or planning. In this paper, we present a novel approach to vehicle localization in dense semantic maps, including vectorized high-definition maps or 3D meshes, using semantic segmentation from a monocular camera. We formulate the localization task as a direct image alignment problem on semantic images, which allows our approach to robustly track the vehicle pose in semantically labeled maps by aligning virtual camera views rendered from the map to sequences of semantically segmented camera images. In contrast to existing visual localization approaches, the system does not require additional keypoint features, handcrafted localization landmark extractors or expensive LiDAR sensors. We demonstrate the wide applicability of our method on a diverse set of semantic mesh maps generated from stereo or LiDAR as well as manually annotated HD maps and show that it achieves reliable and accurate localization in real-time.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge