ProtT3: Protein-to-Text Generation for Text-based Protein Understanding
Paper and Code
May 21, 2024
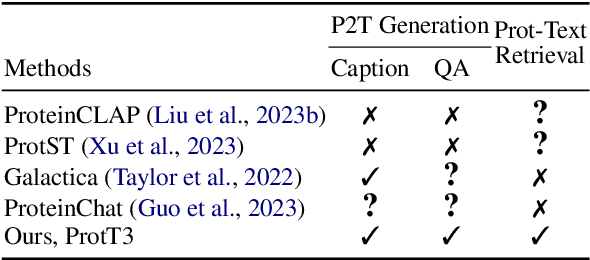

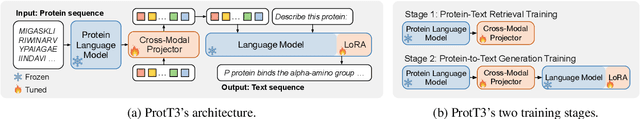
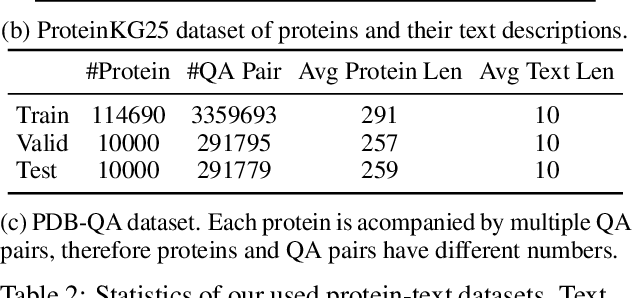
Language Models (LMs) excel in understanding textual descriptions of proteins, as evident in biomedical question-answering tasks. However, their capability falters with raw protein data, such as amino acid sequences, due to a deficit in pretraining on such data. Conversely, Protein Language Models (PLMs) can understand and convert protein data into high-quality representations, but struggle to process texts. To address their limitations, we introduce ProtT3, a framework for Protein-to-Text Generation for Text-based Protein Understanding. ProtT3 empowers an LM to understand protein sequences of amino acids by incorporating a PLM as its protein understanding module, enabling effective protein-to-text generation. This collaboration between PLM and LM is facilitated by a cross-modal projector (i.e., Q-Former) that bridges the modality gap between the PLM's representation space and the LM's input space. Unlike previous studies focusing on protein property prediction and protein-text retrieval, we delve into the largely unexplored field of protein-to-text generation. To facilitate comprehensive benchmarks and promote future research, we establish quantitative evaluations for protein-text modeling tasks, including protein captioning, protein question-answering, and protein-text retrieval. Our experiments show that ProtT3 substantially surpasses current baselines, with ablation studies further highlighting the efficacy of its core components. Our code is available at https://github.com/acharkq/ProtT3.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge