Functional Geometry Guided Protein Sequence and Backbone Structure Co-Design
Paper and Code
Oct 09, 2023
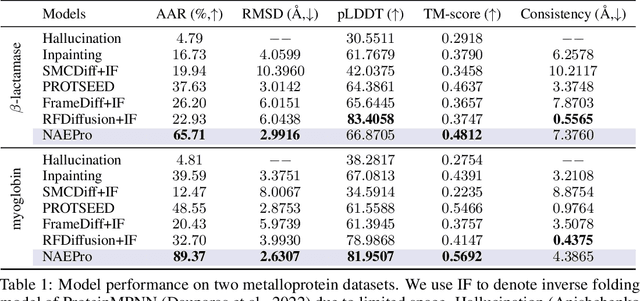
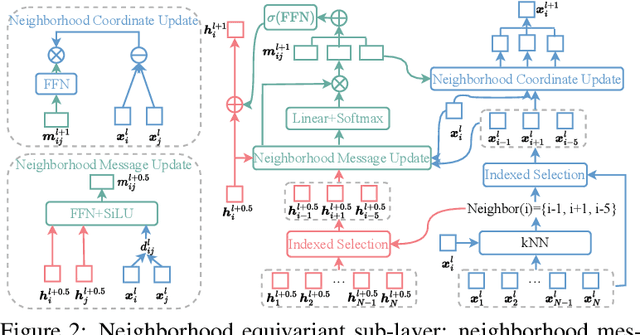
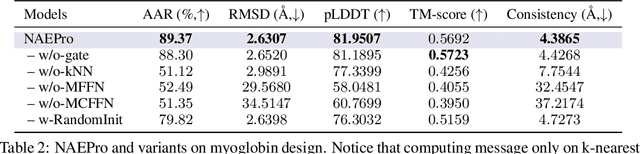
Proteins are macromolecules responsible for essential functions in almost all living organisms. Designing reasonable proteins with desired functions is crucial. A protein's sequence and structure are strongly correlated and they together determine its function. In this paper, we propose NAEPro, a model to jointly design Protein sequence and structure based on automatically detected functional sites. NAEPro is powered by an interleaving network of attention and equivariant layers, which can capture global correlation in a whole sequence and local influence from nearest amino acids in three dimensional (3D) space. Such an architecture facilitates effective yet economic message passing at two levels. We evaluate our model and several strong baselines on two protein datasets, $\beta$-lactamase and myoglobin. Experimental results show that our model consistently achieves the highest amino acid recovery rate, TM-score, and the lowest RMSD among all competitors. These findings prove the capability of our model to design protein sequences and structures that closely resemble their natural counterparts. Furthermore, in-depth analysis further confirms our model's ability to generate highly effective proteins capable of binding to their target metallocofactors. We provide code, data and models in Github.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge