Extracting Interpretable Logic Rules from Graph Neural Networks
Paper and Code
Mar 25, 2025
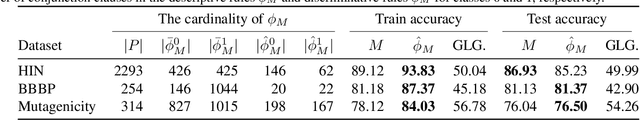
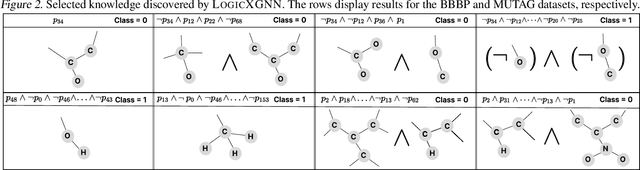
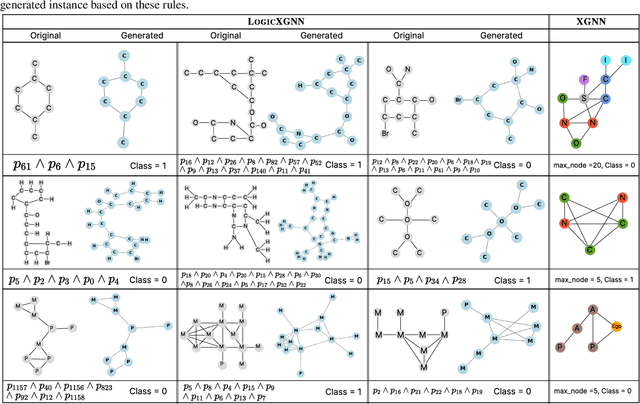
Graph neural networks (GNNs) operate over both input feature spaces and combinatorial graph structures, making it challenging to understand the rationale behind their predictions. As GNNs gain widespread popularity and demonstrate success across various domains, such as drug discovery, studying their interpretability has become a critical task. To address this, many explainability methods have been proposed, with recent efforts shifting from instance-specific explanations to global concept-based explainability. However, these approaches face several limitations, such as relying on predefined concepts and explaining only a limited set of patterns. To address this, we propose a novel framework, LOGICXGNN, for extracting interpretable logic rules from GNNs. LOGICXGNN is model-agnostic, efficient, and data-driven, eliminating the need for predefined concepts. More importantly, it can serve as a rule-based classifier and even outperform the original neural models. Its interpretability facilitates knowledge discovery, as demonstrated by its ability to extract detailed and accurate chemistry knowledge that is often overlooked by existing methods. Another key advantage of LOGICXGNN is its ability to generate new graph instances in a controlled and transparent manner, offering significant potential for applications such as drug design. We empirically demonstrate these merits through experiments on real-world datasets such as MUTAG and BBBP.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge