Efficient Data Selection for Training Genomic Perturbation Models
Paper and Code
Mar 18, 2025
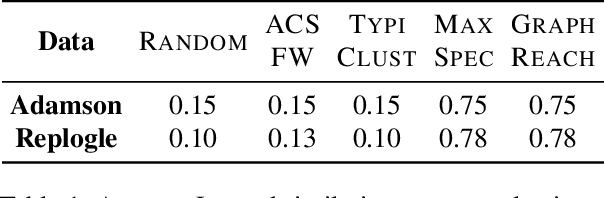
Genomic studies, including CRISPR-based PerturbSeq analyses, face a vast hypothesis space, while gene perturbations remain costly and time-consuming. Gene expression models based on graph neural networks are trained to predict the outcomes of gene perturbations to facilitate such experiments. Active learning methods are often employed to train these models due to the cost of the genomic experiments required to build the training set. However, poor model initialization in active learning can result in suboptimal early selections, wasting time and valuable resources. While typical active learning mitigates this issue over many iterations, the limited number of experimental cycles in genomic studies exacerbates the risk. To this end, we propose graph-based one-shot data selection methods for training gene expression models. Unlike active learning, one-shot data selection predefines the gene perturbations before training, hence removing the initialization bias. The data selection is motivated by theoretical studies of graph neural network generalization. The criteria are defined over the input graph and are optimized with submodular maximization. We compare them empirically to baselines and active learning methods that are state-of-the-art on this problem. The results demonstrate that graph-based one-shot data selection achieves comparable accuracy while alleviating the aforementioned risks.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge