Controllability, Multiplexing, and Transfer Learning in Networks using Evolutionary Learning
Paper and Code
Nov 14, 2018
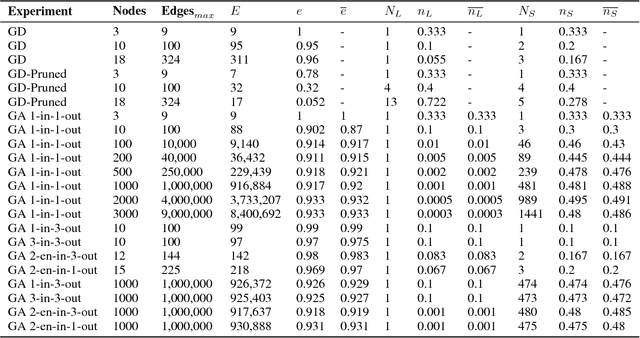
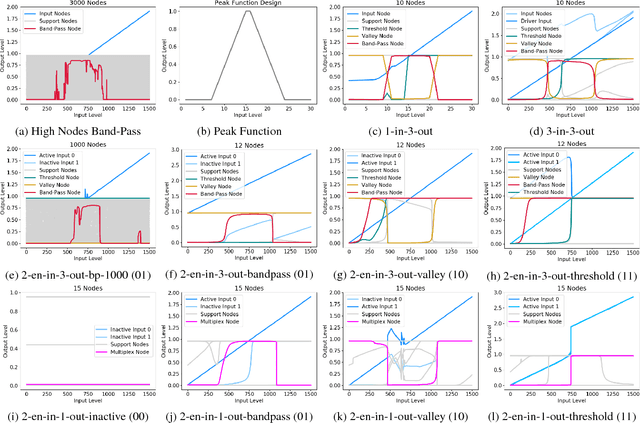

Networks are fundamental building blocks for representing data, and computations. Remarkable progress in learning in structurally defined (shallow or deep) networks has recently been achieved. Here we introduce evolutionary exploratory search and learning method of topologically flexible networks under the constraint of producing elementary computational steady-state input-output operations. Our results include; (1) the identification of networks, over four orders of magnitude, implementing computation of steady-state input-output functions, such as a band-pass filter, a threshold function, and an inverse band-pass function. Next, (2) the learned networks are technically controllable as only a small number of driver nodes are required to move the system to a new state. Furthermore, we find that the fraction of required driver nodes is constant during evolutionary learning, suggesting a stable system design. (3), our framework allows multiplexing of different computations using the same network. For example, using a binary representation of the inputs, the network can readily compute three different input-output functions. Finally, (4) the proposed evolutionary learning demonstrates transfer learning. If the system learns one function A, then learning B requires on average less number of steps as compared to learning B from tabula rasa. We conclude that the constrained evolutionary learning produces large robust controllable circuits, capable of multiplexing and transfer learning. Our study suggests that network-based computations of steady-state functions, representing either cellular modules of cell-to-cell communication networks or internal molecular circuits communicating within a cell, could be a powerful model for biologically inspired computing. This complements conceptualizations such as attractor based models, or reservoir computing.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge