A New Ensemble Learning Framework for 3D Biomedical Image Segmentation
Paper and Code
Dec 10, 2018
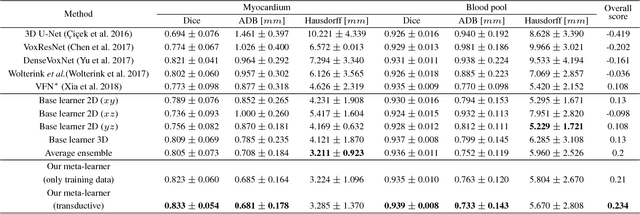
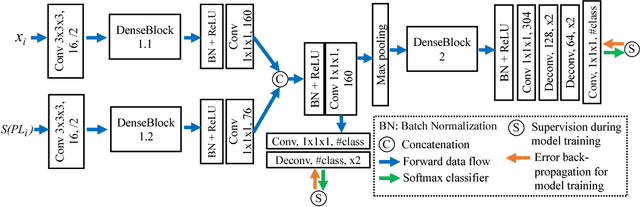
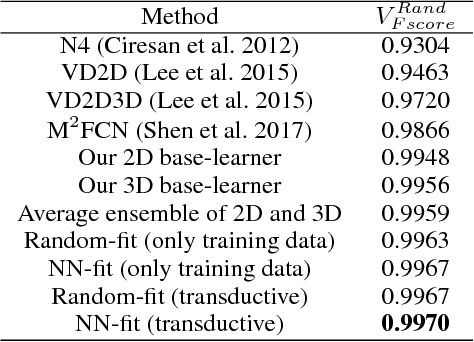
3D image segmentation plays an important role in biomedical image analysis. Many 2D and 3D deep learning models have achieved state-of-the-art segmentation performance on 3D biomedical image datasets. Yet, 2D and 3D models have their own strengths and weaknesses, and by unifying them together, one may be able to achieve more accurate results. In this paper, we propose a new ensemble learning framework for 3D biomedical image segmentation that combines the merits of 2D and 3D models. First, we develop a fully convolutional network based meta-learner to learn how to improve the results from 2D and 3D models (base-learners). Then, to minimize over-fitting for our sophisticated meta-learner, we devise a new training method that uses the results of the base-learners as multiple versions of "ground truths". Furthermore, since our new meta-learner training scheme does not depend on manual annotation, it can utilize abundant unlabeled 3D image data to further improve the model. Extensive experiments on two public datasets (the HVSMR 2016 Challenge dataset and the mouse piriform cortex dataset) show that our approach is effective under fully-supervised, semi-supervised, and transductive settings, and attains superior performance over state-of-the-art image segmentation methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge