A Bottom-up Approach for Pancreas Segmentation using Cascaded Superpixels and (Deep) Image Patch Labeling
Paper and Code
Mar 07, 2016
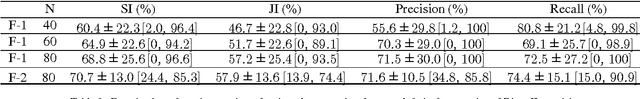



Robust automated organ segmentation is a prerequisite for computer-aided diagnosis (CAD), quantitative imaging analysis and surgical assistance. For high-variability organs such as the pancreas, previous approaches report undesirably low accuracies. We present a bottom-up approach for pancreas segmentation in abdominal CT scans that is based on a hierarchy of information propagation by classifying image patches at different resolutions; and cascading superpixels. There are four stages: 1) decomposing CT slice images as a set of disjoint boundary-preserving superpixels; 2) computing pancreas class probability maps via dense patch labeling; 3) classifying superpixels by pooling both intensity and probability features to form empirical statistics in cascaded random forest frameworks; and 4) simple connectivity based post-processing. The dense image patch labeling are conducted by: efficient random forest classifier on image histogram, location and texture features; and more expensive (but with better specificity) deep convolutional neural network classification on larger image windows (with more spatial contexts). Evaluation of the approach is performed on a database of 80 manually segmented CT volumes in six-fold cross-validation (CV). Our achieved results are comparable, or better than the state-of-the-art methods (evaluated by "leave-one-patient-out"), with Dice 70.7% and Jaccard 57.9%. The computational efficiency has been drastically improved in the order of 6~8 minutes, comparing with others of ~10 hours per case. Finally, we implement a multi-atlas label fusion (MALF) approach for pancreas segmentation using the same datasets. Under six-fold CV, our bottom-up segmentation method significantly outperforms its MALF counterpart: (70.7 +/- 13.0%) versus (52.5 +/- 20.8%) in Dice. Deep CNN patch labeling confidences offer more numerical stability, reflected by smaller standard deviations.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge