A Benchmark for End-to-End Zero-Shot Biomedical Relation Extraction with LLMs: Experiments with OpenAI Models
Paper and Code
Apr 05, 2025
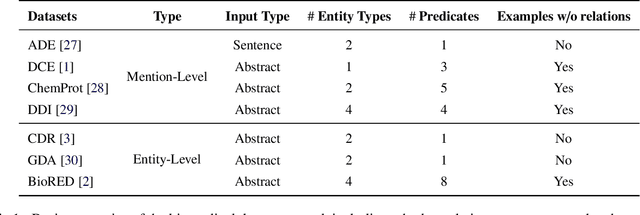
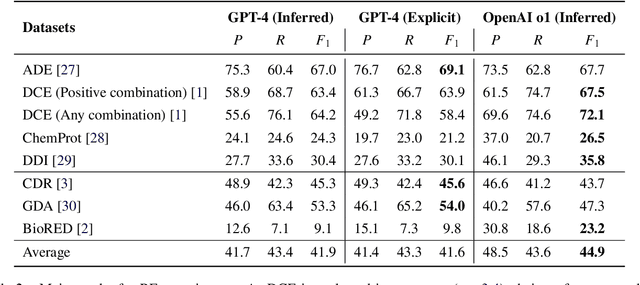
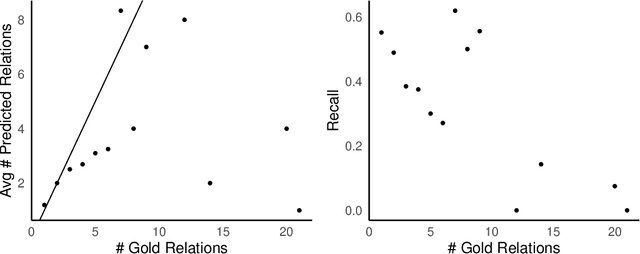
Objective: Zero-shot methodology promises to cut down on costs of dataset annotation and domain expertise needed to make use of NLP. Generative large language models trained to align with human goals have achieved high zero-shot performance across a wide variety of tasks. As of yet, it is unclear how well these models perform on biomedical relation extraction (RE). To address this knowledge gap, we explore patterns in the performance of OpenAI LLMs across a diverse sampling of RE tasks. Methods: We use OpenAI GPT-4-turbo and their reasoning model o1 to conduct end-to-end RE experiments on seven datasets. We use the JSON generation capabilities of GPT models to generate structured output in two ways: (1) by defining an explicit schema describing the structure of relations, and (2) using a setting that infers the structure from the prompt language. Results: Our work is the first to study and compare the performance of the GPT-4 and o1 for the end-to-end zero-shot biomedical RE task across a broad array of datasets. We found the zero-shot performances to be proximal to that of fine-tuned methods. The limitations of this approach are that it performs poorly on instances containing many relations and errs on the boundaries of textual mentions. Conclusion: Recent large language models exhibit promising zero-shot capabilities in complex biomedical RE tasks, offering competitive performance with reduced dataset curation and NLP modeling needs at the cost of increased computing, potentially increasing medical community accessibility. Addressing the limitations we identify could further boost reliability. The code, data, and prompts for all our experiments are publicly available: https://github.com/bionlproc/ZeroShotRE
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge