3D Denoisers are Good 2D Teachers: Molecular Pretraining via Denoising and Cross-Modal Distillation
Paper and Code
Sep 08, 2023
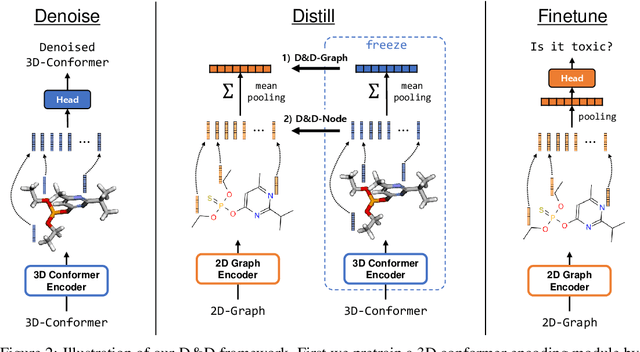
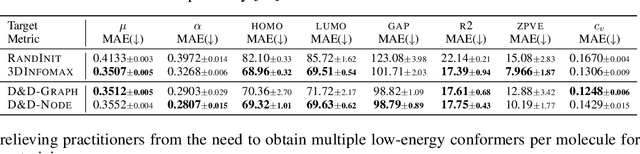
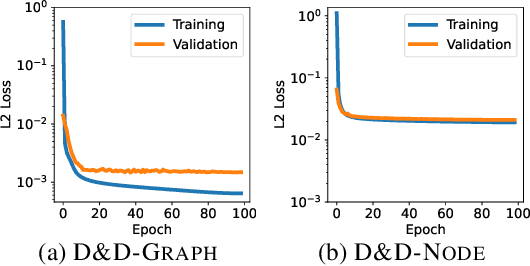
Pretraining molecular representations from large unlabeled data is essential for molecular property prediction due to the high cost of obtaining ground-truth labels. While there exist various 2D graph-based molecular pretraining approaches, these methods struggle to show statistically significant gains in predictive performance. Recent work have thus instead proposed 3D conformer-based pretraining under the task of denoising, which led to promising results. During downstream finetuning, however, models trained with 3D conformers require accurate atom-coordinates of previously unseen molecules, which are computationally expensive to acquire at scale. In light of this limitation, we propose D&D, a self-supervised molecular representation learning framework that pretrains a 2D graph encoder by distilling representations from a 3D denoiser. With denoising followed by cross-modal knowledge distillation, our approach enjoys use of knowledge obtained from denoising as well as painless application to downstream tasks with no access to accurate conformers. Experiments on real-world molecular property prediction datasets show that the graph encoder trained via D&D can infer 3D information based on the 2D graph and shows superior performance and label-efficiency against other baselines.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge