Yang Young Lu
Interactive Training: Feedback-Driven Neural Network Optimization
Oct 02, 2025Abstract:Traditional neural network training typically follows fixed, predefined optimization recipes, lacking the flexibility to dynamically respond to instabilities or emerging training issues. In this paper, we introduce Interactive Training, an open-source framework that enables real-time, feedback-driven intervention during neural network training by human experts or automated AI agents. At its core, Interactive Training uses a control server to mediate communication between users or agents and the ongoing training process, allowing users to dynamically adjust optimizer hyperparameters, training data, and model checkpoints. Through three case studies, we demonstrate that Interactive Training achieves superior training stability, reduced sensitivity to initial hyperparameters, and improved adaptability to evolving user needs, paving the way toward a future training paradigm where AI agents autonomously monitor training logs, proactively resolve instabilities, and optimize training dynamics.
Error-controlled non-additive interaction discovery in machine learning models
Aug 30, 2024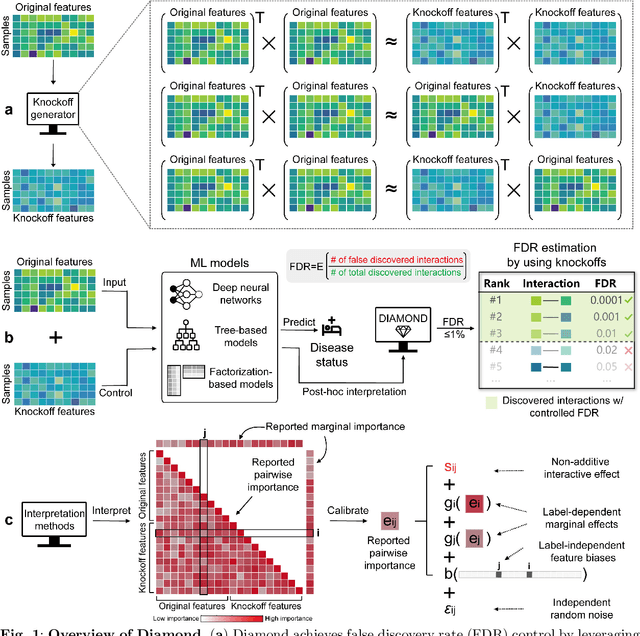
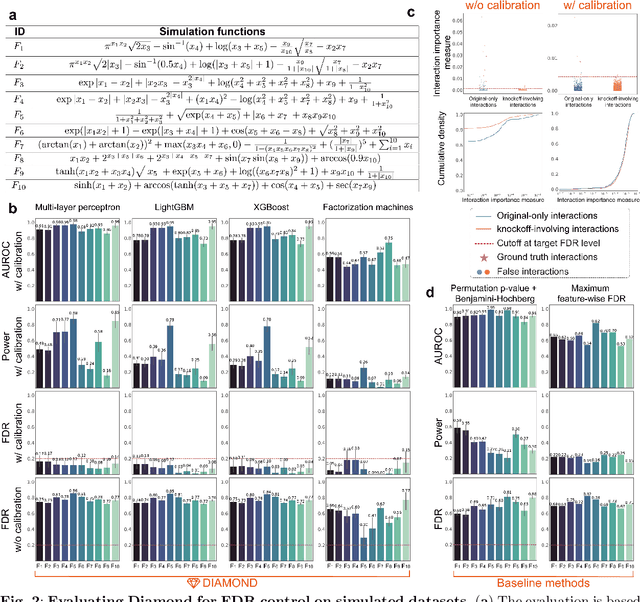
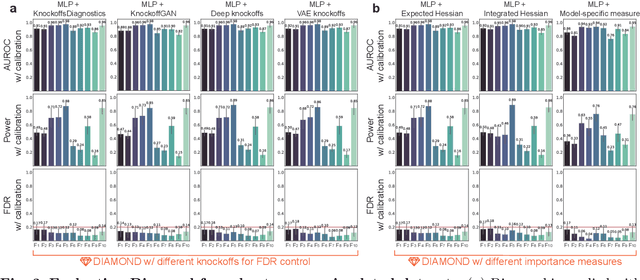
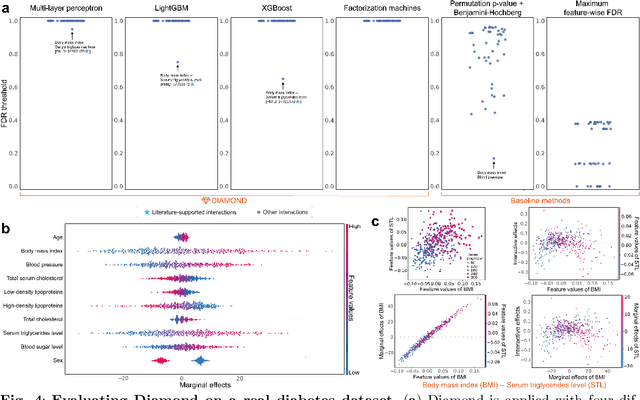
Abstract:Machine learning (ML) models are powerful tools for detecting complex patterns within data, yet their "black box" nature limits their interpretability, hindering their use in critical domains like healthcare and finance. To address this challenge, interpretable ML methods have been developed to explain how features influence model predictions. However, these methods often focus on univariate feature importance, overlooking the complex interactions between features that ML models are capable of capturing. Recognizing this limitation, recent efforts have aimed to extend these methods to discover feature interactions, but existing approaches struggle with robustness and error control, especially under data perturbations. In this study, we introduce Diamond, a novel method for trustworthy feature interaction discovery. Diamond uniquely integrates the model-X knockoffs framework to control the false discovery rate (FDR), ensuring that the proportion of falsely discovered interactions remains low. We further address the challenges of using off-the-shelf interaction importance measures by proposing a calibration procedure that refines these measures to maintain the desired FDR. Diamond's applicability spans a wide range of ML models, including deep neural networks, tree-based models, and factorization-based models. Our empirical evaluations on both simulated and real datasets across various biomedical studies demonstrate Diamond's utility in enabling more reliable data-driven scientific discoveries. This method represents a significant step forward in the deployment of ML models for scientific innovation and hypothesis generation.
DeepROCK: Error-controlled interaction detection in deep neural networks
Sep 26, 2023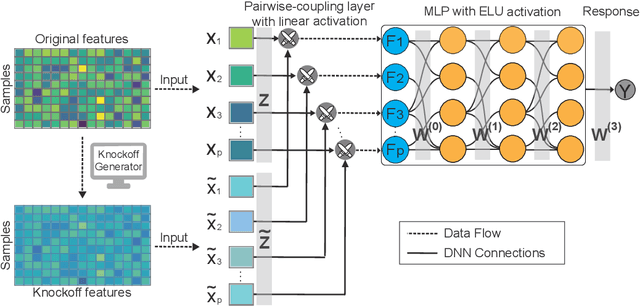
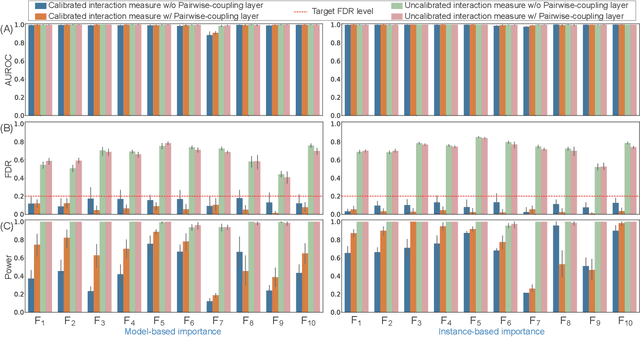
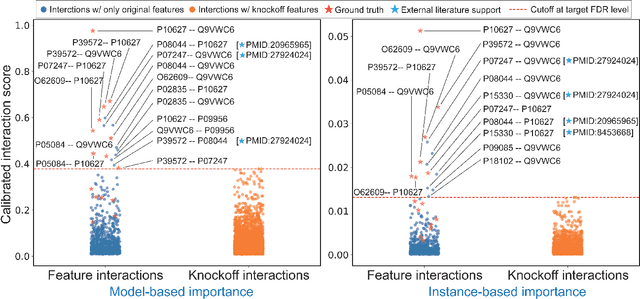
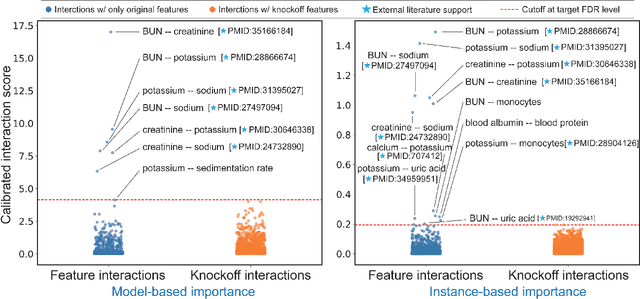
Abstract:The complexity of deep neural networks (DNNs) makes them powerful but also makes them challenging to interpret, hindering their applicability in error-intolerant domains. Existing methods attempt to reason about the internal mechanism of DNNs by identifying feature interactions that influence prediction outcomes. However, such methods typically lack a systematic strategy to prioritize interactions while controlling confidence levels, making them difficult to apply in practice for scientific discovery and hypothesis validation. In this paper, we introduce a method, called DeepROCK, to address this limitation by using knockoffs, which are dummy variables that are designed to mimic the dependence structure of a given set of features while being conditionally independent of the response. Together with a novel DNN architecture involving a pairwise-coupling layer, DeepROCK jointly controls the false discovery rate (FDR) and maximizes statistical power. In addition, we identify a challenge in correctly controlling FDR using off-the-shelf feature interaction importance measures. DeepROCK overcomes this challenge by proposing a calibration procedure applied to existing interaction importance measures to make the FDR under control at a target level. Finally, we validate the effectiveness of DeepROCK through extensive experiments on simulated and real datasets.
DeepPINK: reproducible feature selection in deep neural networks
Sep 06, 2018

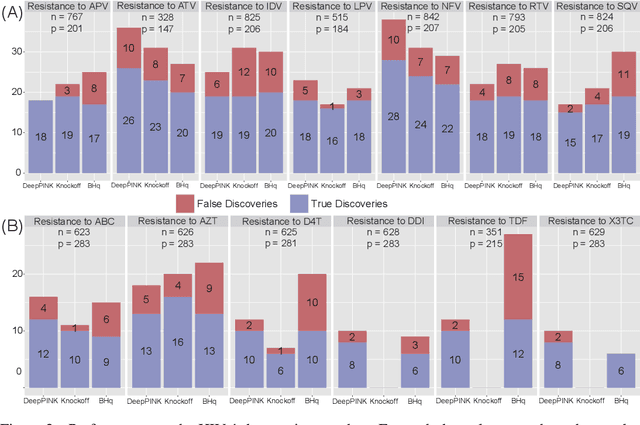

Abstract:Deep learning has become increasingly popular in both supervised and unsupervised machine learning thanks to its outstanding empirical performance. However, because of their intrinsic complexity, most deep learning methods are largely treated as black box tools with little interpretability. Even though recent attempts have been made to facilitate the interpretability of deep neural networks (DNNs), existing methods are susceptible to noise and lack of robustness. Therefore, scientists are justifiably cautious about the reproducibility of the discoveries, which is often related to the interpretability of the underlying statistical models. In this paper, we describe a method to increase the interpretability and reproducibility of DNNs by incorporating the idea of feature selection with controlled error rate. By designing a new DNN architecture and integrating it with the recently proposed knockoffs framework, we perform feature selection with a controlled error rate, while maintaining high power. This new method, DeepPINK (Deep feature selection using Paired-Input Nonlinear Knockoffs), is applied to both simulated and real data sets to demonstrate its empirical utility.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge