Xinyuan Shao
Memorizing SAM: 3D Medical Segment Anything Model with Memorizing Transformer
Dec 18, 2024Abstract:Segment Anything Models (SAMs) have gained increasing attention in medical image analysis due to their zero-shot generalization capability in segmenting objects of unseen classes and domains when provided with appropriate user prompts. Addressing this performance gap is important to fully leverage the pre-trained weights of SAMs, particularly in the domain of volumetric medical image segmentation, where accuracy is important but well-annotated 3D medical data for fine-tuning is limited. In this work, we investigate whether introducing the memory mechanism as a plug-in, specifically the ability to memorize and recall internal representations of past inputs, can improve the performance of SAM with limited computation cost. To this end, we propose Memorizing SAM, a novel 3D SAM architecture incorporating a memory Transformer as a plug-in. Unlike conventional memorizing Transformers that save the internal representation during training or inference, our Memorizing SAM utilizes existing highly accurate internal representation as the memory source to ensure the quality of memory. We evaluate the performance of Memorizing SAM in 33 categories from the TotalSegmentator dataset, which indicates that Memorizing SAM can outperform state-of-the-art 3D SAM variant i.e., FastSAM3D with an average Dice increase of 11.36% at the cost of only 4.38 millisecond increase in inference time. The source code is publicly available at https://github.com/swedfr/memorizingSAM
Performance and Non-adversarial Robustness of the Segment Anything Model 2 in Surgical Video Segmentation
Aug 07, 2024Abstract:Fully supervised deep learning (DL) models for surgical video segmentation have been shown to struggle with non-adversarial, real-world corruptions of image quality including smoke, bleeding, and low illumination. Foundation models for image segmentation, such as the segment anything model (SAM) that focuses on interactive prompt-based segmentation, move away from semantic classes and thus can be trained on larger and more diverse data, which offers outstanding zero-shot generalization with appropriate user prompts. Recently, building upon this success, SAM-2 has been proposed to further extend the zero-shot interactive segmentation capabilities from independent frame-by-frame to video segmentation. In this paper, we present a first experimental study evaluating SAM-2's performance on surgical video data. Leveraging the SegSTRONG-C MICCAI EndoVIS 2024 sub-challenge dataset, we assess SAM-2's effectiveness on uncorrupted endoscopic sequences and evaluate its non-adversarial robustness on videos with corrupted image quality simulating smoke, bleeding, and low brightness conditions under various prompt strategies. Our experiments demonstrate that SAM-2, in zero-shot manner, can achieve competitive or even superior performance compared to fully-supervised deep learning models on surgical video data, including under non-adversarial corruptions of image quality. Additionally, SAM-2 consistently outperforms the original SAM and its medical variants across all conditions. Finally, frame-sparse prompting can consistently outperform frame-wise prompting for SAM-2, suggesting that allowing SAM-2 to leverage its temporal modeling capabilities leads to more coherent and accurate segmentation compared to frequent prompting.
FastSAM-3DSlicer: A 3D-Slicer Extension for 3D Volumetric Segment Anything Model with Uncertainty Quantification
Jul 17, 2024Abstract:Accurate segmentation of anatomical structures and pathological regions in medical images is crucial for diagnosis, treatment planning, and disease monitoring. While the Segment Anything Model (SAM) and its variants have demonstrated impressive interactive segmentation capabilities on image types not seen during training without the need for domain adaptation or retraining, their practical application in volumetric 3D medical imaging workflows has been hindered by the lack of a user-friendly interface. To address this challenge, we introduce FastSAM-3DSlicer, a 3D Slicer extension that integrates both 2D and 3D SAM models, including SAM-Med2D, MedSAM, SAM-Med3D, and FastSAM-3D. Building on the well-established open-source 3D Slicer platform, our extension enables efficient, real-time segmentation of 3D volumetric medical images, with seamless interaction and visualization. By automating the handling of raw image data, user prompts, and segmented masks, FastSAM-3DSlicer provides a streamlined, user-friendly interface that can be easily incorporated into medical image analysis workflows. Performance evaluations reveal that the FastSAM-3DSlicer extension running FastSAM-3D achieves low inference times of only 1.09 seconds per volume on CPU and 0.73 seconds per volume on GPU, making it well-suited for real-time interactive segmentation. Moreover, we introduce an uncertainty quantification scheme that leverages the rapid inference capabilities of FastSAM-3D for practical implementation, further enhancing its reliability and applicability in medical settings. FastSAM-3DSlicer offers an interactive platform and user interface for 2D and 3D interactive volumetric medical image segmentation, offering a powerful combination of efficiency, precision, and ease of use with SAMs. The source code and a video demonstration are publicly available at https://github.com/arcadelab/FastSAM3D_slicer.
FastSAM3D: An Efficient Segment Anything Model for 3D Volumetric Medical Images
Mar 14, 2024
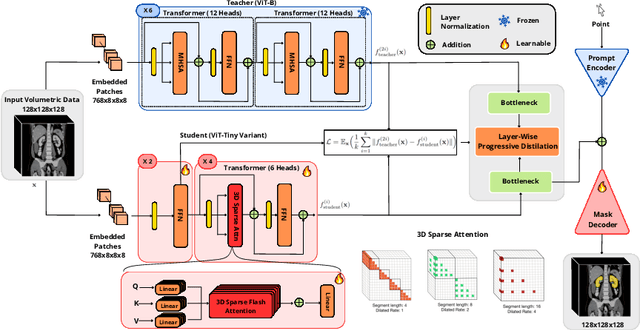


Abstract:Segment anything models (SAMs) are gaining attention for their zero-shot generalization capability in segmenting objects of unseen classes and in unseen domains when properly prompted. Interactivity is a key strength of SAMs, allowing users to iteratively provide prompts that specify objects of interest to refine outputs. However, to realize the interactive use of SAMs for 3D medical imaging tasks, rapid inference times are necessary. High memory requirements and long processing delays remain constraints that hinder the adoption of SAMs for this purpose. Specifically, while 2D SAMs applied to 3D volumes contend with repetitive computation to process all slices independently, 3D SAMs suffer from an exponential increase in model parameters and FLOPS. To address these challenges, we present FastSAM3D which accelerates SAM inference to 8 milliseconds per 128*128*128 3D volumetric image on an NVIDIA A100 GPU. This speedup is accomplished through 1) a novel layer-wise progressive distillation scheme that enables knowledge transfer from a complex 12-layer ViT-B to a lightweight 6-layer ViT-Tiny variant encoder without training from scratch; and 2) a novel 3D sparse flash attention to replace vanilla attention operators, substantially reducing memory needs and improving parallelization. Experiments on three diverse datasets reveal that FastSAM3D achieves a remarkable speedup of 527.38x compared to 2D SAMs and 8.75x compared to 3D SAMs on the same volumes without significant performance decline. Thus, FastSAM3D opens the door for low-cost truly interactive SAM-based 3D medical imaging segmentation with commonly used GPU hardware. Code is available at https://github.com/arcadelab/FastSAM3D.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge