Wenbiao Zhou
Beijing StoneWise Technology Co Ltd
Lingo3DMol: Generation of a Pocket-based 3D Molecule using a Language Model
May 17, 2023Abstract:Structure-based drug design powered by deep generative models have attracted increasing research interest in recent years. Language models have demonstrated a robust capacity for generating valid molecules in 2D structures, while methods based on geometric deep learning can directly produce molecules with accurate 3D coordinates. Inspired by both methods, this article proposes a pocket-based 3D molecule generation method that leverages the language model with the ability to generate 3D coordinates. High quality protein-ligand complex data are insufficient; hence, a perturbation and restoration pre-training task is designed that can utilize vast amounts of small-molecule data. A new molecular representation, a fragment-based SMILES with local and global coordinates, is also presented, enabling the language model to learn molecular topological structures and spatial position information effectively. Ultimately, CrossDocked and DUD-E dataset is employed for evaluation and additional metrics are introduced. This method achieves state-of-the-art performance in nearly all metrics, notably in terms of binding patterns, drug-like properties, rational conformations, and inference speed. Our model is available as an online service to academic users via sw3dmg.stonewise.cn
G2GT: Retrosynthesis Prediction with Graph to Graph Attention Neural Network and Self-Training
Apr 19, 2022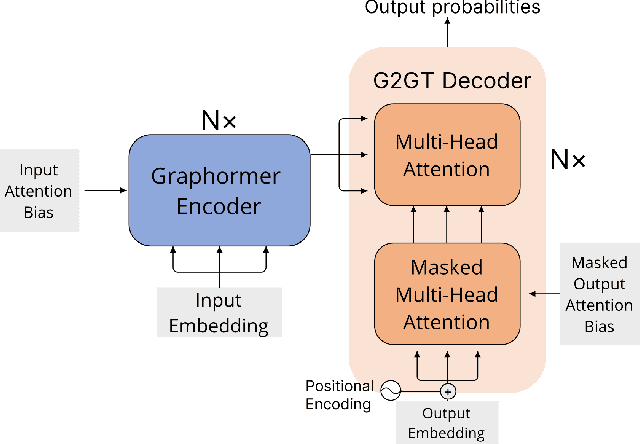
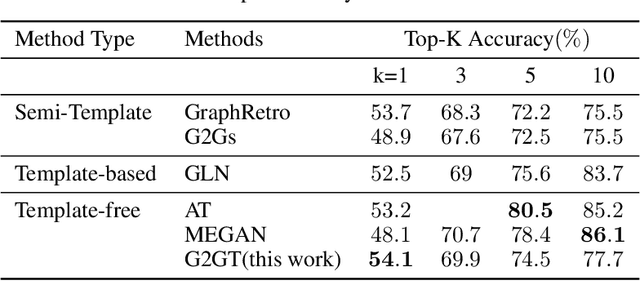
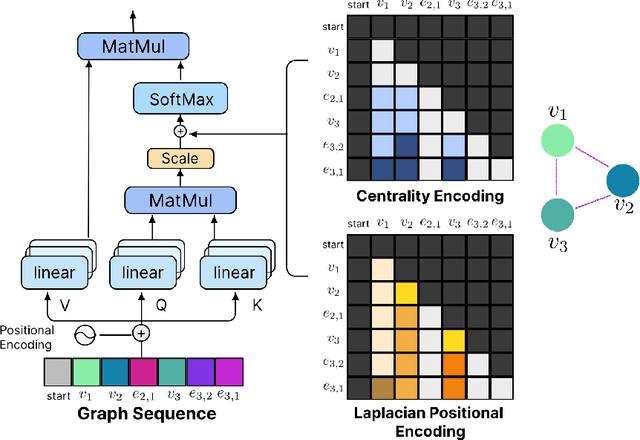
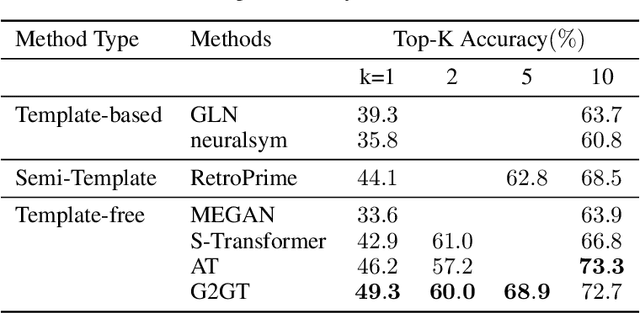
Abstract:Retrosynthesis prediction is one of the fundamental challenges in organic chemistry and related fields. The goal is to find reactants molecules that can synthesize product molecules. To solve this task, we propose a new graph-to-graph transformation model, G2GT, in which the graph encoder and graph decoder are built upon the standard transformer structure. We also show that self-training, a powerful data augmentation method that utilizes unlabeled molecule data, can significantly improve the model's performance. Inspired by the reaction type label and ensemble learning, we proposed a novel weak ensemble method to enhance diversity. We combined beam search, nucleus, and top-k sampling methods to further improve inference diversity and proposed a simple ranking algorithm to retrieve the final top-10 results. We achieved new state-of-the-art results on both the USPTO-50K dataset, with top1 accuracy of 54%, and the larger data set USPTO-full, with top1 accuracy of 50%, and competitive top-10 results.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge