Vincent Jugnon
BN-SCAFFOLD: controlling the drift of Batch Normalization statistics in Federated Learning
Oct 04, 2024
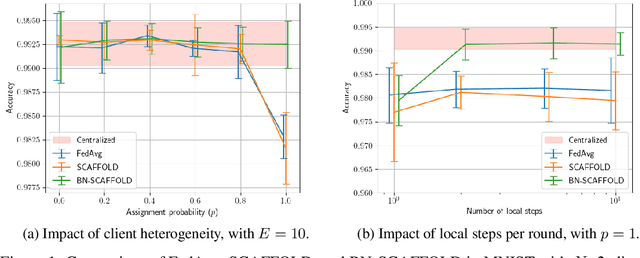
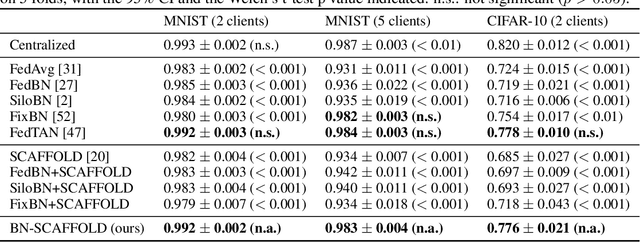
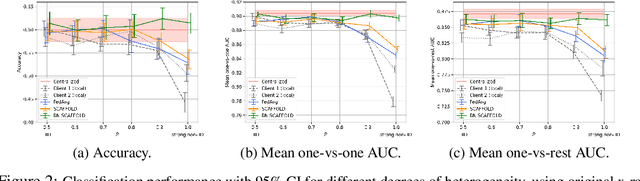
Abstract:Federated Learning (FL) is gaining traction as a learning paradigm for training Machine Learning (ML) models in a decentralized way. Batch Normalization (BN) is ubiquitous in Deep Neural Networks (DNN), as it improves convergence and generalization. However, BN has been reported to hinder performance of DNNs in heterogeneous FL. Recently, the FedTAN algorithm has been proposed to mitigate the effect of heterogeneity on BN, by aggregating BN statistics and gradients from all the clients. However, it has a high communication cost, that increases linearly with the depth of the DNN. SCAFFOLD is a variance reduction algorithm, that estimates and corrects the client drift in a communication-efficient manner. Despite its promising results in heterogeneous FL settings, it has been reported to underperform for models with BN. In this work, we seek to revive SCAFFOLD, and more generally variance reduction, as an efficient way of training DNN with BN in heterogeneous FL. We introduce a unified theoretical framework for analyzing the convergence of variance reduction algorithms in the BN-DNN setting, inspired of by the work of Wang et al. 2023, and show that SCAFFOLD is unable to remove the bias introduced by BN. We thus propose the BN-SCAFFOLD algorithm, which extends the client drift correction of SCAFFOLD to BN statistics. We prove convergence using the aforementioned framework and validate the theoretical results with experiments on MNIST and CIFAR-10. BN-SCAFFOLD equals the performance of FedTAN, without its high communication cost, outperforming Federated Averaging (FedAvg), SCAFFOLD, and other FL algorithms designed to mitigate BN heterogeneity.
How to Register a Live onto a Liver ? Partial Matching in the Space of Varifolds
Apr 12, 2022
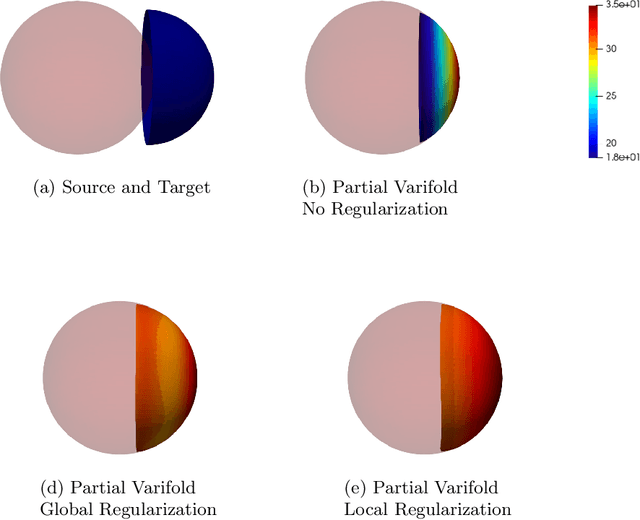



Abstract:Partial shapes correspondences is a problem that often occurs in computer vision (occlusion, evolution in time...). In medical imaging, data may come from different modalities and be acquired under different conditions which leads to variations in shapes and topologies. In this paper we use an asymmetric data dissimilarity term applicable to various geometric shapes like sets of curves or surfaces, assessing the embedding of a shape into another one without relying on correspondences. It is designed as a data attachment for the Large Deformation Diffeomorphic Metric Mapping (LDDMM) framework, allowing to compute a meaningful deformation of one shape onto a subset of the other. We refine it in order to control the resulting non-rigid deformations and provide consistent deformations of the shapes along with their ambient space. We show that partial matching can be used for robust multi-modal liver registration between a Computed Tomography (CT) volume and a Cone Beam Computed Tomography (CBCT) volume. The 3D imaging of the patient CBCT at point of care that we call live is truncated while the CT pre-intervention provides a full visualization of the liver. The proposed method allows the truncated surfaces from CBCT to be aligned non-rigidly, yet realistically, with surfaces from CT with an average distance of 2.6mm(+/- 2.2). The generated deformations extend consistently to the liver volume, and are evaluated on points of interest for the physicians, with an average distance of 5.8mm (+/- 2.7) for vessels bifurcations and 5.13mm (+/- 2.5) for tumors landmarks. Such multi-modality volumes registrations would help the physicians in the perspective of navigating their tools in the patient's anatomy to locate structures that are hardly visible in the CBCT used during their procedures. Our code is available at https://github.com/plantonsanti/PartialMatchingVarifolds.
* 30 pages, 11 figures, Special Issue: Information Processing in Medical Imaging (IPMI) 2021, Accepted for publication at the Journal of Machine Learning for Biomedical Imaging (MELBA) https://www.melba-journal.org
Partial Matching in the Space of Varifolds
Mar 23, 2021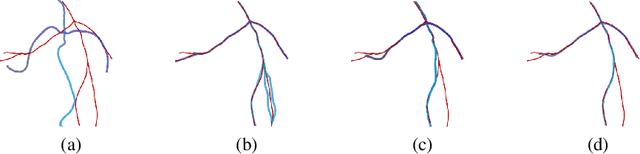


Abstract:In computer vision and medical imaging, the problem of matching structures finds numerous applications from automatic annotation to data reconstruction. The data however, while corresponding to the same anatomy, are often very different in topology or shape and might only partially match each other. We introduce a new asymmetric data dissimilarity term for various geometric shapes like sets of curves or surfaces. This term is based on the Varifold shape representation and assesses the embedding of a shape into another one without relying on correspondences between points. It is designed as data attachment for the Large Deformation Diffeomorphic Metric Mapping (LDDMM) framework, allowing to compute meaningful deformation of one shape onto a subset of the other. Registrations are illustrated on sets of synthetic 3D curves, real vascular trees and livers' surfaces from two different modalities: Computed Tomography (CT) and Cone Beam Computed Tomography (CBCT). All experiments show that this data dissimilarity term leads to coherent partial matching despite the topological differences.
Database Annotation with few Examples: An Atlas-based Framework using Diffeomorphic Registration of 3D Trees
Sep 25, 2020


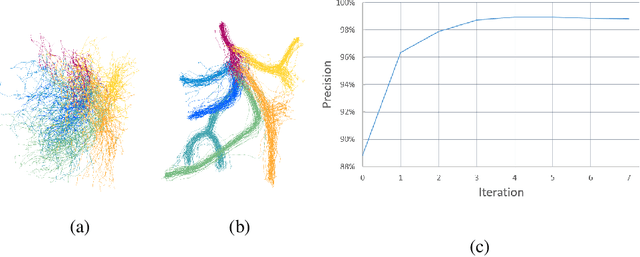
Abstract:Automatic annotation of anatomical structures can help simplify workflow during interventions in numerous clinical applications but usually involves a large amount of annotated data. The complexity of the labeling task, together with the lack of representative data, slows down the development of robust solutions. In this paper, we propose a solution requiring very few annotated cases to label 3D pelvic arterial trees of patients with benign prostatic hyperplasia. We take advantage of Large Deformation Diffeomorphic Metric Mapping (LDDMM) to perform registration based on meaningful deformations from which we build an atlas. Branch pairing is then computed from the atlas to new cases using optimal transport to ensure one-to-one correspondence during the labeling process. To tackle topological variations in the tree, which usually degrades the performance of atlas-based techniques, we propose a simple bottom-up label assignment adapted to the pelvic anatomy. The proposed method achieves 97.6\% labeling precision with only 5 cases for training, while in comparison learning-based methods only reach 82.2\% on such small training sets.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge