Tomomi Nobashi
CT organ segmentation using GPU data augmentation, unsupervised labels and IOU loss
Nov 27, 2018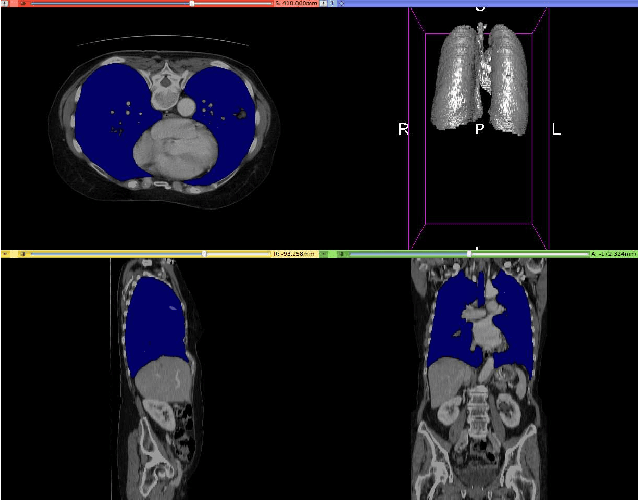

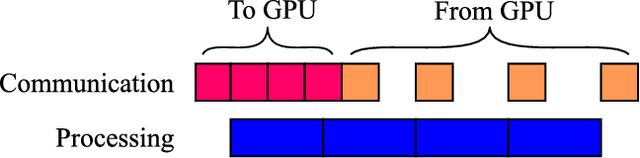
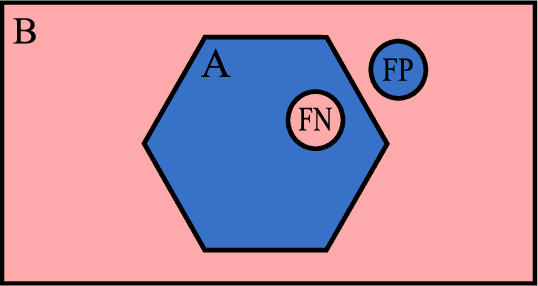
Abstract:Fully-convolutional neural networks have achieved superior performance in a variety of image segmentation tasks. However, their training requires laborious manual annotation of large datasets, as well as acceleration by parallel processors with high-bandwidth memory, such as GPUs. We show that simple models can achieve competitive accuracy for organ segmentation on CT images when trained with extensive data augmentation, which leverages existing graphics hardware to quickly apply geometric and photometric transformations to 3D image data. On 3 mm^3 CT volumes, our GPU implementation is 2.6-8X faster than a widely-used CPU version, including communication overhead. We also show how to automatically generate training labels using rudimentary morphological operations, which are efficiently computed by 3D Fourier transforms. We combined fully-automatic labels for the lungs and bone with semi-automatic ones for the liver, kidneys and bladder, to create a dataset of 130 labeled CT scans. To achieve the best results from data augmentation, our model uses the intersection-over-union (IOU) loss function, a close relative of the Dice loss. We discuss its mathematical properties and explain why it outperforms the usual weighted cross-entropy loss for unbalanced segmentation tasks. We conclude that there is no unique IOU loss function, as the naive one belongs to a broad family of functions with the same essential properties. When combining data augmentation with the IOU loss, our model achieves a Dice score of 78-92% for each organ. The trained model, code and dataset will be made publicly available, to further medical imaging research.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge