Stéphane Robin
Exact and approximate inference in graphical models: variable elimination and beyond
Mar 12, 2018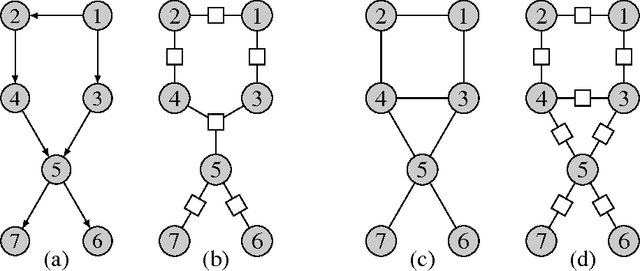

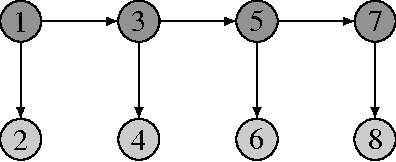
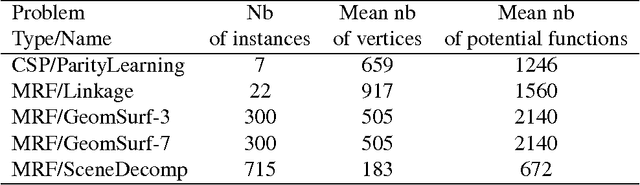
Abstract:Probabilistic graphical models offer a powerful framework to account for the dependence structure between variables, which is represented as a graph. However, the dependence between variables may render inference tasks intractable. In this paper we review techniques exploiting the graph structure for exact inference, borrowed from optimisation and computer science. They are built on the principle of variable elimination whose complexity is dictated in an intricate way by the order in which variables are eliminated. The so-called treewidth of the graph characterises this algorithmic complexity: low-treewidth graphs can be processed efficiently. The first message that we illustrate is therefore the idea that for inference in graphical model, the number of variables is not the limiting factor, and it is worth checking for the treewidth before turning to approximate methods. We show how algorithms providing an upper bound of the treewidth can be exploited to derive a 'good' elimination order enabling to perform exact inference. The second message is that when the treewidth is too large, algorithms for approximate inference linked to the principle of variable elimination, such as loopy belief propagation and variational approaches, can lead to accurate results while being much less time consuming than Monte-Carlo approaches. We illustrate the techniques reviewed in this article on benchmarks of inference problems in genetic linkage analysis and computer vision, as well as on hidden variables restoration in coupled Hidden Markov Models.
A closed-form approach to Bayesian inference in tree-structured graphical models
May 01, 2017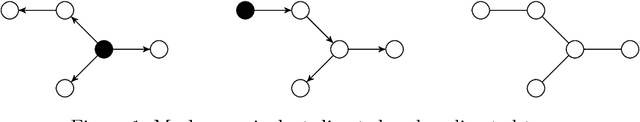

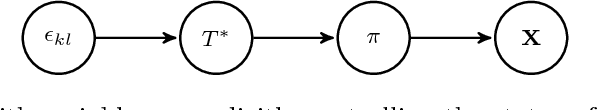
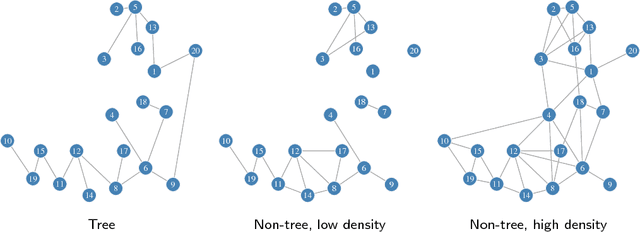
Abstract:We consider the inference of the structure of an undirected graphical model in an exact Bayesian framework. More specifically we aim at achieving the inference with close-form posteriors, avoiding any sampling step. This task would be intractable without any restriction on the considered graphs, so we limit our exploration to mixtures of spanning trees. We consider the inference of the structure of an undirected graphical model in a Bayesian framework. To avoid convergence issues and highly demanding Monte Carlo sampling, we focus on exact inference. More specifically we aim at achieving the inference with close-form posteriors, avoiding any sampling step. To this aim, we restrict the set of considered graphs to mixtures of spanning trees. We investigate under which conditions on the priors - on both tree structures and parameters - exact Bayesian inference can be achieved. Under these conditions, we derive a fast an exact algorithm to compute the posterior probability for an edge to belong to {the tree model} using an algebraic result called the Matrix-Tree theorem. We show that the assumption we have made does not prevent our approach to perform well on synthetic and flow cytometry data.
Exact Bayesian inference for off-line change-point detection in tree-structured graphical models
Jun 16, 2016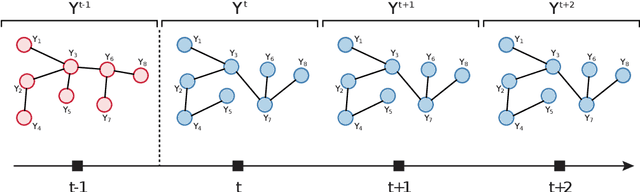
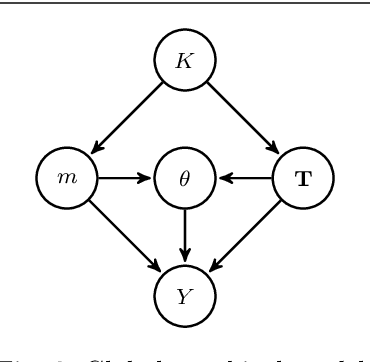
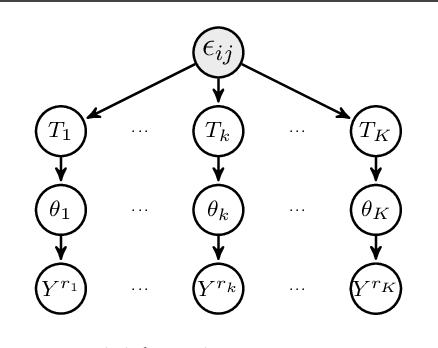
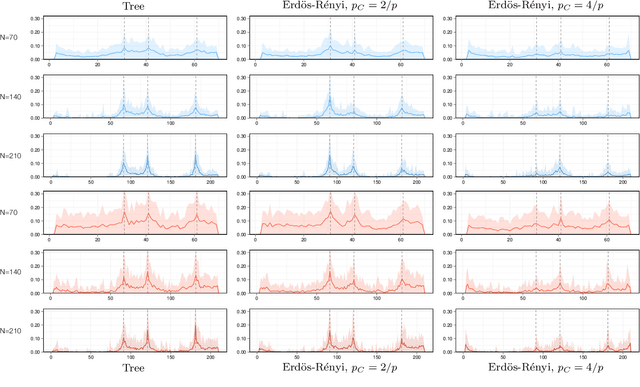
Abstract:We consider the problem of change-point detection in multivariate time-series. The multivariate distribution of the observations is supposed to follow a graphical model, whose graph and parameters are affected by abrupt changes throughout time. We demonstrate that it is possible to perform exact Bayesian inference whenever one considers a simple class of undirected graphs called spanning trees as possible structures. We are then able to integrate on the graph and segmentation spaces at the same time by combining classical dynamic programming with algebraic results pertaining to spanning trees. In particular, we show that quantities such as posterior distributions for change-points or posterior edge probabilities over time can efficiently be obtained. We illustrate our results on both synthetic and experimental data arising from biology and neuroscience.
Hidden Markov Models with mixtures as emission distributions
Jun 22, 2012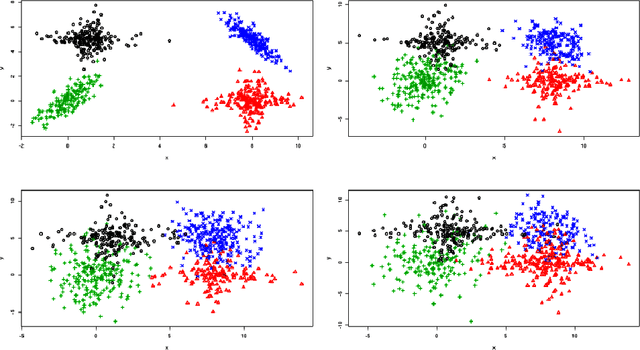
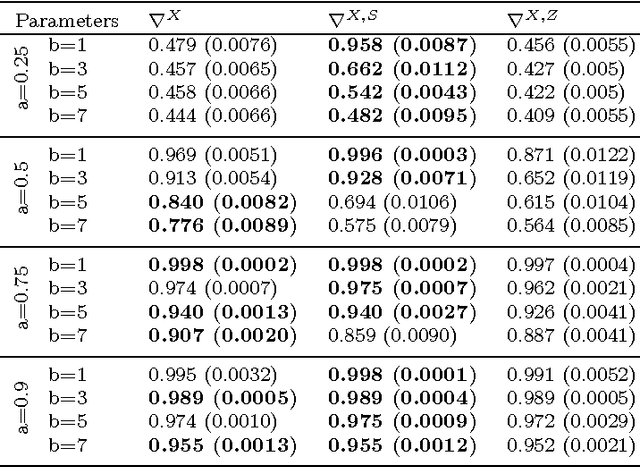
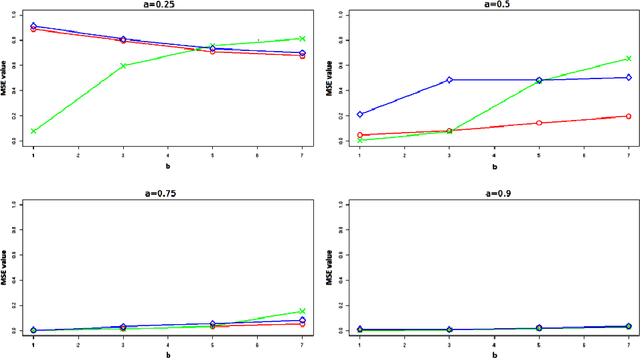
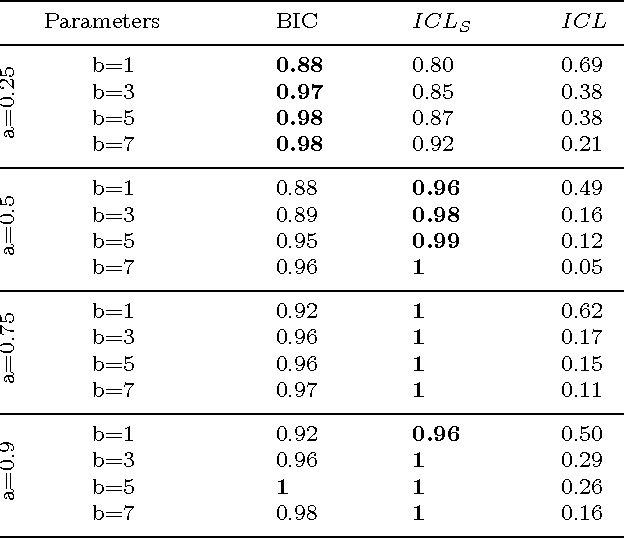
Abstract:In unsupervised classification, Hidden Markov Models (HMM) are used to account for a neighborhood structure between observations. The emission distributions are often supposed to belong to some parametric family. In this paper, a semiparametric modeling where the emission distributions are a mixture of parametric distributions is proposed to get a higher flexibility. We show that the classical EM algorithm can be adapted to infer the model parameters. For the initialisation step, starting from a large number of components, a hierarchical method to combine them into the hidden states is proposed. Three likelihood-based criteria to select the components to be combined are discussed. To estimate the number of hidden states, BIC-like criteria are derived. A simulation study is carried out both to determine the best combination between the merging criteria and the model selection criteria and to evaluate the accuracy of classification. The proposed method is also illustrated using a biological dataset from the model plant Arabidopsis thaliana. A R package HMMmix is freely available on the CRAN.
Variational Bayes approach for model aggregation in unsupervised classification with Markovian dependency
May 04, 2011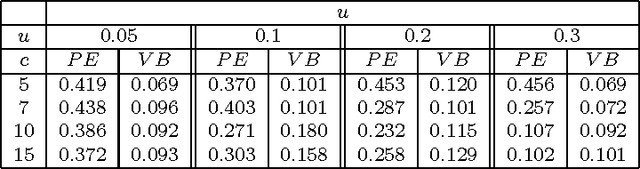
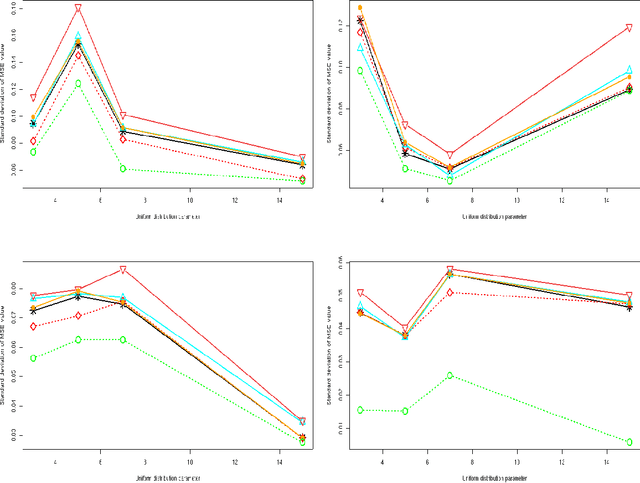
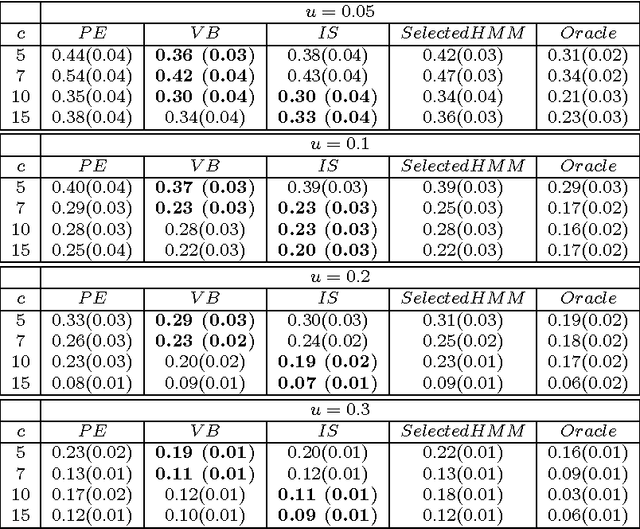
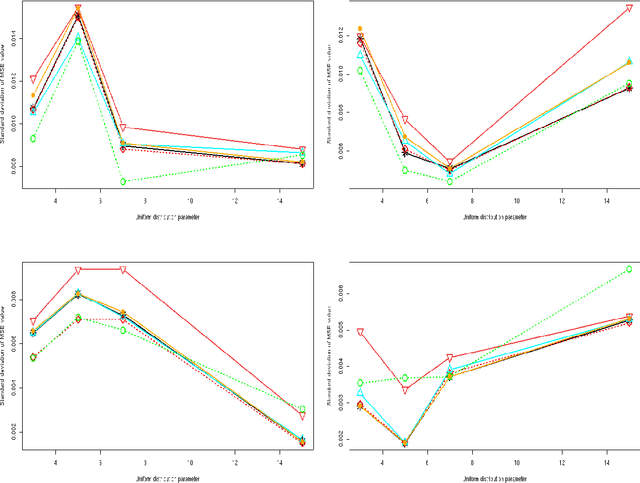
Abstract:We consider a binary unsupervised classification problem where each observation is associated with an unobserved label that we want to retrieve. More precisely, we assume that there are two groups of observation: normal and abnormal. The `normal' observations are coming from a known distribution whereas the distribution of the `abnormal' observations is unknown. Several models have been developed to fit this unknown distribution. In this paper, we propose an alternative based on a mixture of Gaussian distributions. The inference is done within a variational Bayesian framework and our aim is to infer the posterior probability of belonging to the class of interest. To this end, it makes no sense to estimate the mixture component number since each mixture model provides more or less relevant information to the posterior probability estimation. By computing a weighted average (named aggregated estimator) over the model collection, Bayesian Model Averaging (BMA) is one way of combining models in order to account for information provided by each model. The aim is then the estimation of the weights and the posterior probability for one specific model. In this work, we derive optimal approximations of these quantities from the variational theory and propose other approximations of the weights. To perform our method, we consider that the data are dependent (Markovian dependency) and hence we consider a Hidden Markov Model. A simulation study is carried out to evaluate the accuracy of the estimates in terms of classification. We also present an application to the analysis of public health surveillance systems.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge