Sir Michael Brady
SVRDA: A Web-based Dataset Annotation Tool for Slice-to-Volume Registration
Nov 27, 2023
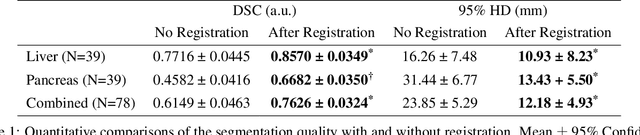


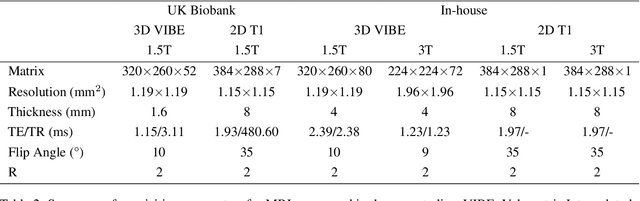
Abstract:Background and Objective: The lack of benchmark datasets has impeded the development of slice-to-volume registration algorithms. Such datasets are difficult to annotate, primarily due to the dimensional difference within data and the dearth of task-specific software. We aim to develop a user-friendly tool to streamline dataset annotation for slice-to-volume registration. Methods: The proposed tool, named SVRDA, is an installation-free web application for platform-agnostic collaborative dataset annotation. It enables efficient transformation manipulation via keyboard shortcuts and smooth case transitions with auto-saving. SVRDA supports configuration-based data loading and adheres to the separation of concerns, offering great flexibility and extensibility for future research. Various supplementary features have been implemented to facilitate slice-to-volume registration. Results: We validated the effectiveness of SVRDA by indirectly evaluating the post-registration segmentation quality on UK Biobank data, observing a dramatic overall improvement (24.02% in the Dice Similarity Coefficient and 48.93% in the 95th percentile Hausdorff distance, respectively) supported by highly statistically significant evidence ($p<0.001$).We further showcased the clinical usage of SVRDA by integrating it into test-retest T1 quantification on in-house magnetic resonance images, leading to more consistent results after registration. Conclusions: SVRDA can facilitate collaborative annotation of benchmark datasets while being potentially applicable to other pipelines incorporating slice-to-volume registration. Full source code and documentation are available at https://github.com/Roldbach/SVRDA
Pieces-of-parts for supervoxel segmentation with global context: Application to DCE-MRI tumour delineation
Apr 18, 2016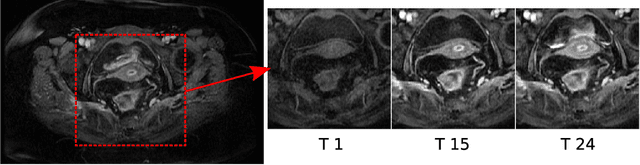

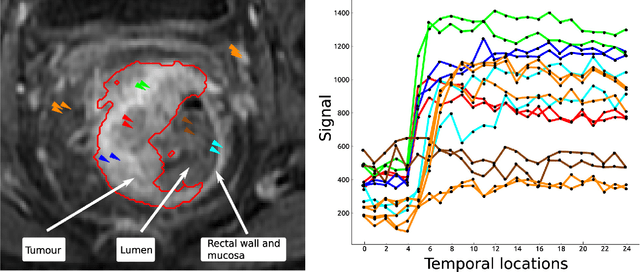
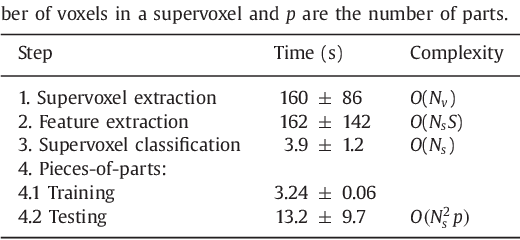
Abstract:Rectal tumour segmentation in dynamic contrast-enhanced MRI (DCE-MRI) is a challenging task, and an automated and consistent method would be highly desirable to improve the modelling and prediction of patient outcomes from tissue contrast enhancement characteristics - particularly in routine clinical practice. A framework is developed to automate DCE-MRI tumour segmentation, by introducing: perfusion-supervoxels to over-segment and classify DCE-MRI volumes using the dynamic contrast enhancement characteristics; and the pieces-of-parts graphical model, which adds global (anatomic) constraints that further refine the supervoxel components that comprise the tumour. The framework was evaluated on 23 DCE-MRI scans of patients with rectal adenocarcinomas, and achieved a voxelwise area-under the receiver operating characteristic curve (AUC) of 0.97 compared to expert delineations. Creating a binary tumour segmentation, 21 of the 23 cases were segmented correctly with a median Dice similarity coefficient (DSC) of 0.63, which is close to the inter-rater variability of this challenging task. A sec- ond study is also included to demonstrate the method's generalisability and achieved a DSC of 0.71. The framework achieves promising results for the underexplored area of rectal tumour segmentation in DCE-MRI, and the methods have potential to be applied to other DCE-MRI and supervoxel segmentation problems
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge