Ruben Verhagen
Supervised Tractogram Filtering using Geometric Deep Learning
Dec 06, 2022



Abstract:A tractogram is a virtual representation of the brain white matter. It is composed of millions of virtual fibers, encoded as 3D polylines, which approximate the white matter axonal pathways. To date, tractograms are the most accurate white matter representation and thus are used for tasks like presurgical planning and investigations of neuroplasticity, brain disorders, or brain networks. However, it is a well-known issue that a large portion of tractogram fibers is not anatomically plausible and can be considered artifacts of the tracking procedure. With Verifyber, we tackle the problem of filtering out such non-plausible fibers using a novel fully-supervised learning approach. Differently from other approaches based on signal reconstruction and/or brain topology regularization, we guide our method with the existing anatomical knowledge of the white matter. Using tractograms annotated according to anatomical principles, we train our model, Verifyber, to classify fibers as either anatomically plausible or non-plausible. The proposed Verifyber model is an original Geometric Deep Learning method that can deal with variable size fibers, while being invariant to fiber orientation. Our model considers each fiber as a graph of points, and by learning features of the edges between consecutive points via the proposed sequence Edge Convolution, it can capture the underlying anatomical properties. The output filtering results highly accurate and robust across an extensive set of experiments, and fast; with a 12GB GPU, filtering a tractogram of 1M fibers requires less than a minute. Verifyber implementation and trained models are available at https://github.com/FBK-NILab/verifyber.
Tractogram filtering of anatomically non-plausible fibers with geometric deep learning
Mar 24, 2020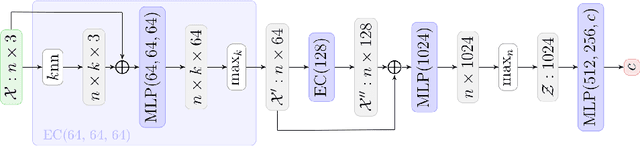
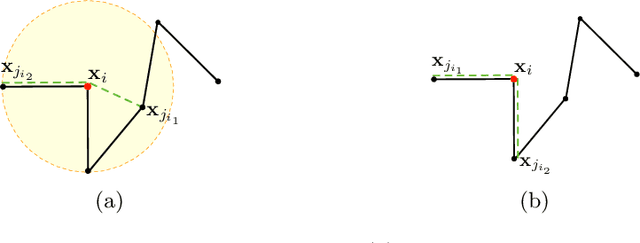
Abstract:Tractograms are virtual representations of the white matter fibers of the brain. They are of primary interest for tasks like presurgical planning, and investigation of neuroplasticity or brain disorders. Each tractogram is composed of millions of fibers encoded as 3D polylines. Unfortunately, a large portion of those fibers are not anatomically plausible and can be considered artifacts of the tracking algorithms. Common methods for tractogram filtering are based on signal reconstruction, a principled approach, but unable to consider the knowledge of brain anatomy. In this work, we address the problem of tractogram filtering as a supervised learning problem by exploiting the ground truth annotations obtained with a recent heuristic method, which labels fibers as either anatomically plausible or non-plausible according to well-established anatomical properties. The intuitive idea is to model a fiber as a point cloud and the goal is to investigate whether and how a geometric deep learning model might capture its anatomical properties. Our contribution is an extension of the Dynamic Edge Convolution model that exploits the sequential relations of points in a fiber and discriminates with high accuracy plausible/non-plausible fibers.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge