Renato Hermoza
Censor-aware Semi-supervised Learning for Survival Time Prediction from Medical Images
May 26, 2022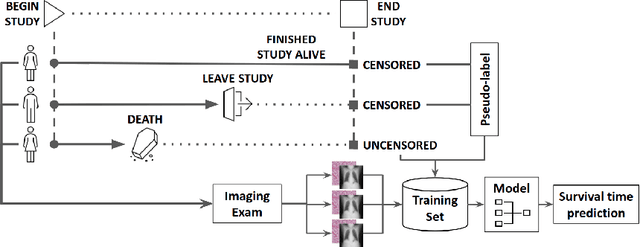
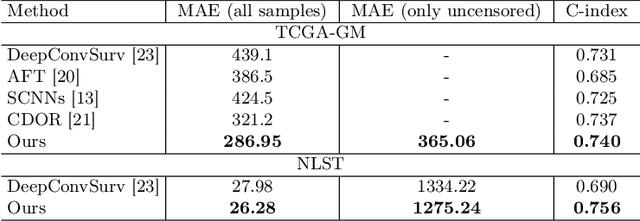
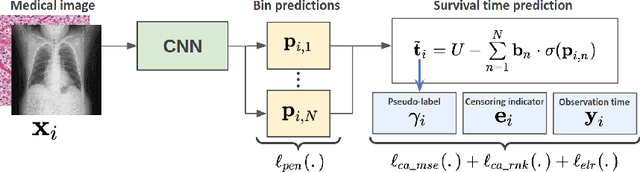
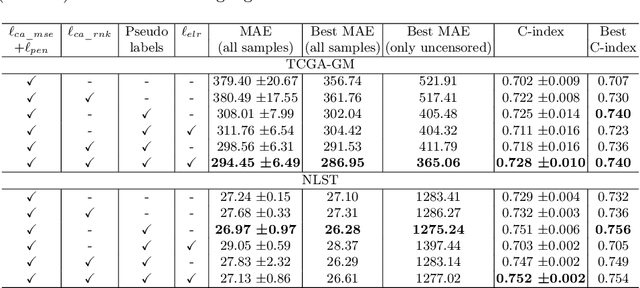
Abstract:Survival time prediction from medical images is important for treatment planning, where accurate estimations can improve healthcare quality. One issue affecting the training of survival models is censored data. Most of the current survival prediction approaches are based on Cox models that can deal with censored data, but their application scope is limited because they output a hazard function instead of a survival time. On the other hand, methods that predict survival time usually ignore censored data, resulting in an under-utilization of the training set. In this work, we propose a new training method that predicts survival time using all censored and uncensored data. We propose to treat censored data as samples with a lower-bound time to death and estimate pseudo labels to semi-supervise a censor-aware survival time regressor. We evaluate our method on pathology and x-ray images from the TCGA-GM and NLST datasets. Our results establish the state-of-the-art survival prediction accuracy on both datasets.
Post-hoc Overall Survival Time Prediction from Brain MRI
Feb 22, 2021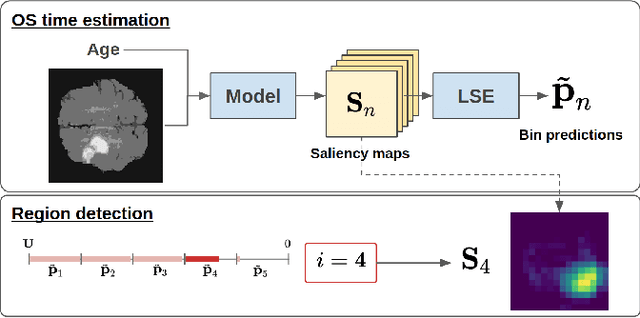
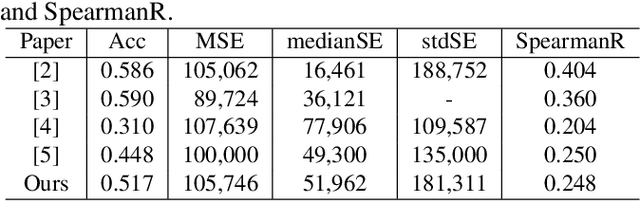
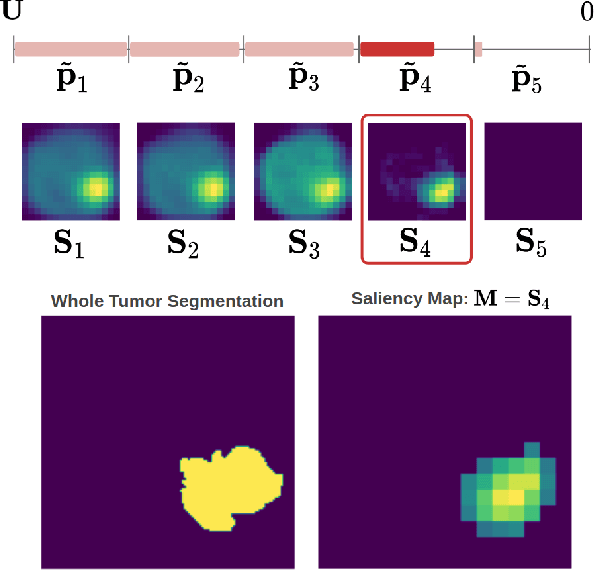

Abstract:Overall survival (OS) time prediction is one of the most common estimates of the prognosis of gliomas and is used to design an appropriate treatment planning. State-of-the-art (SOTA) methods for OS time prediction follow a pre-hoc approach that require computing the segmentation map of the glioma tumor sub-regions (necrotic, edema tumor, enhancing tumor) for estimating OS time. However, the training of the segmentation methods require ground truth segmentation labels which are tedious and expensive to obtain. Given that most of the large-scale data sets available from hospitals are unlikely to contain such precise segmentation, those SOTA methods have limited applicability. In this paper, we introduce a new post-hoc method for OS time prediction that does not require segmentation map annotation for training. Our model uses medical image and patient demographics (represented by age) as inputs to estimate the OS time and to estimate a saliency map that localizes the tumor as a way to explain the OS time prediction in a post-hoc manner. It is worth emphasizing that although our model can localize tumors, it uses only the ground truth OS time as training signal, i.e., no segmentation labels are needed. We evaluate our post-hoc method on the Multimodal Brain Tumor Segmentation Challenge (BraTS) 2019 data set and show that it achieves competitive results compared to pre-hoc methods with the advantage of not requiring segmentation labels for training.
Region Proposals for Saliency Map Refinement for Weakly-supervised Disease Localisation and Classification
May 22, 2020


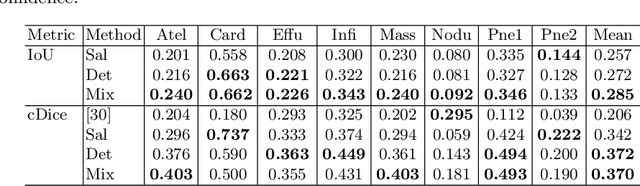
Abstract:The deployment of automated systems to diagnose diseases from medical images is challenged by the requirement to localise the diagnosed diseases to justify or explain the classification decision. This requirement is hard to fulfil because most of the training sets available to develop these systems only contain global annotations, making the localisation of diseases a weakly supervised approach. The main methods designed for weakly supervised disease classification and localisation rely on saliency or attention maps that are not specifically trained for localisation, or on region proposals that can not be refined to produce accurate detections. In this paper, we introduce a new model that combines region proposal and saliency detection to overcome both limitations for weakly supervised disease classification and localisation. Using the ChestX-ray14 data set, we show that our proposed model establishes the new state-of-the-art for weakly-supervised disease diagnosis and localisation.
3D Reconstruction of Incomplete Archaeological Objects Using a Generative Adversarial Network
Mar 10, 2018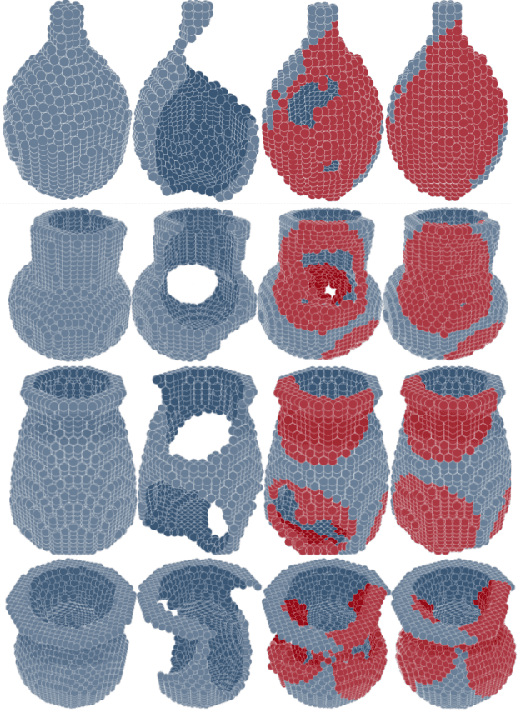

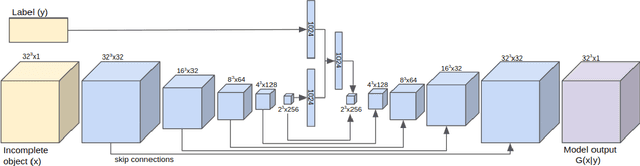
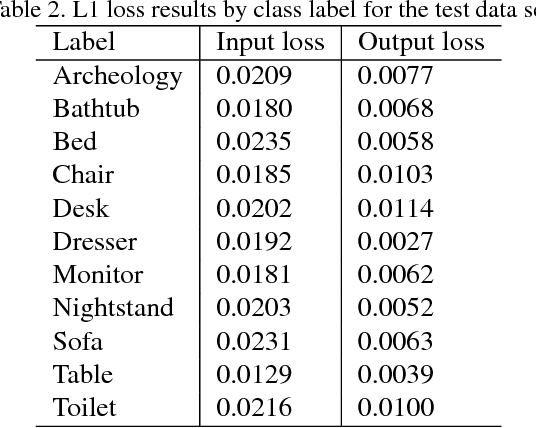
Abstract:We introduce a data-driven approach to aid the repairing and conservation of archaeological objects: ORGAN, an object reconstruction generative adversarial network (GAN). By using an encoder-decoder 3D deep neural network on a GAN architecture, and combining two loss objectives: a completion loss and an Improved Wasserstein GAN loss, we can train a network to effectively predict the missing geometry of damaged objects. As archaeological objects can greatly differ between them, the network is conditioned on a variable, which can be a culture, a region or any metadata of the object. In our results, we show that our method can recover most of the information from damaged objects, even in cases where more than half of the voxels are missing, without producing many errors.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge