Rahul Roy
A novel approach to navigate the taxonomic hierarchy to address the Open-World Scenarios in Medicinal Plant Classification
Feb 24, 2025



Abstract:In this article, we propose a novel approach for plant hierarchical taxonomy classification by posing the problem as an open class problem. It is observed that existing methods for medicinal plant classification often fail to perform hierarchical classification and accurately identifying unknown species, limiting their effectiveness in comprehensive plant taxonomy classification. Thus we address the problem of unknown species classification by assigning it best hierarchical labels. We propose a novel method, which integrates DenseNet121, Multi-Scale Self-Attention (MSSA) and cascaded classifiers for hierarchical classification. The approach systematically categorizes medicinal plants at multiple taxonomic levels, from phylum to species, ensuring detailed and precise classification. Using multi scale space attention, the model captures both local and global contextual information from the images, improving the distinction between similar species and the identification of new ones. It uses attention scores to focus on important features across multiple scales. The proposed method provides a solution for hierarchical classification, showcasing superior performance in identifying both known and unknown species. The model was tested on two state-of-art datasets with and without background artifacts and so that it can be deployed to tackle real word application. We used unknown species for testing our model. For unknown species the model achieved an average accuracy of 83.36%, 78.30%, 60.34% and 43.32% for predicting correct phylum, class, order and family respectively. Our proposed model size is almost four times less than the existing state of the art methods making it easily deploy able in real world application.
Mutual Information and Ensemble Based Feature Recommender for Renal Cancer Stage Classification
Sep 28, 2022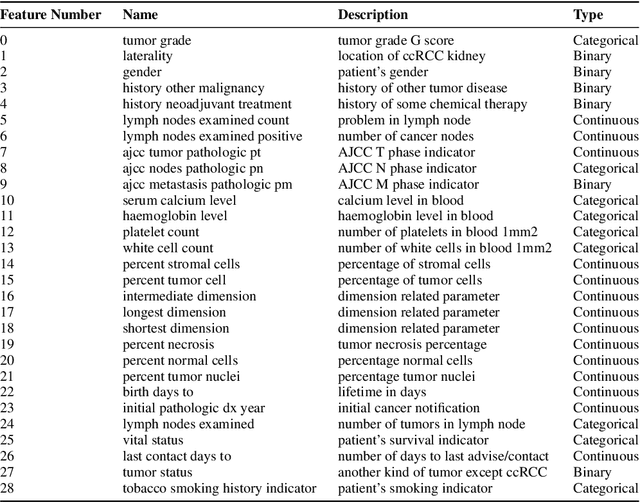
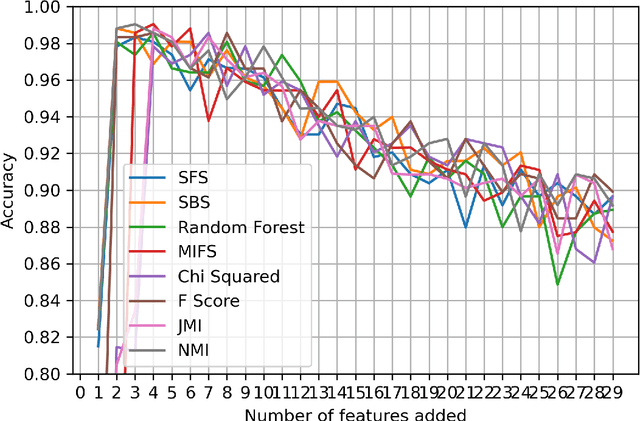
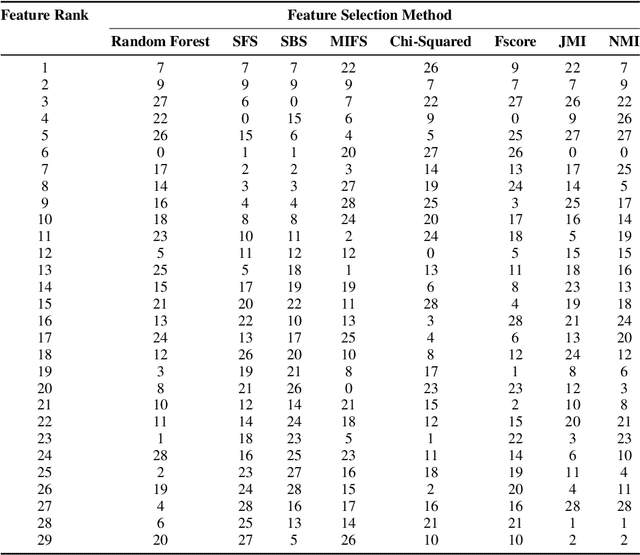
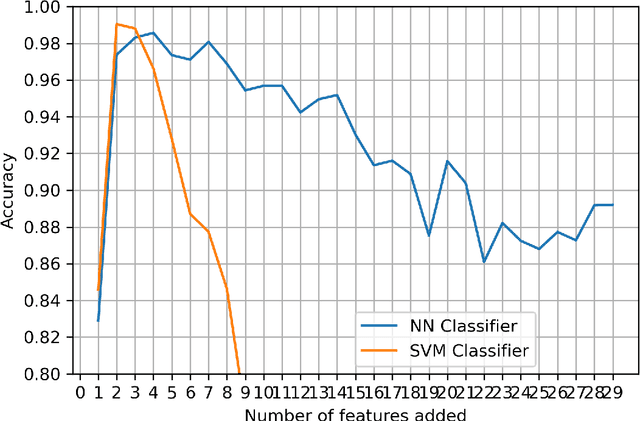
Abstract:Kidney is an essential organ in human body. It maintains homeostasis and removes harmful substances through urine. Renal cell carcinoma (RCC) is the most common form of kidney cancer. Around 90\% of all kidney cancers are attributed to RCC. Most harmful type of RCC is clear cell renal cell carcinoma (ccRCC) that makes up about 80\% of all RCC cases. Early and accurate detection of ccRCC is necessary to prevent further spreading of the disease in other organs. In this article, a detailed experimentation is done to identify important features which can aid in diagnosing ccRCC at different stages. The ccRCC dataset is obtained from The Cancer Genome Atlas (TCGA). A novel mutual information and ensemble based feature ranking approach considering the order of features obtained from 8 popular feature selection methods is proposed. Performance of the proposed method is evaluated by overall classification accuracy obtained using 2 different classifiers (ANN and SVM). Experimental results show that the proposed feature ranking method is able to attain a higher accuracy (96.6\% and 98.6\% using SVM and NN, respectively) for classifying different stages of ccRCC with a reduced feature set as compared to existing work. It is also to be noted that, out of 3 distinguishing features as mentioned by the existing TNM system (proposed by AJCC and UICC), our proposed method was able to select two of them (size of tumour, metastasis status) as the top-most ones. This establishes the efficacy of our proposed approach.
Mixed Uncertainty Sets for Robust Combinatorial Optimization
Jan 21, 2019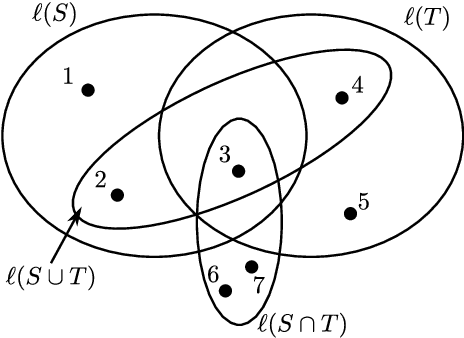
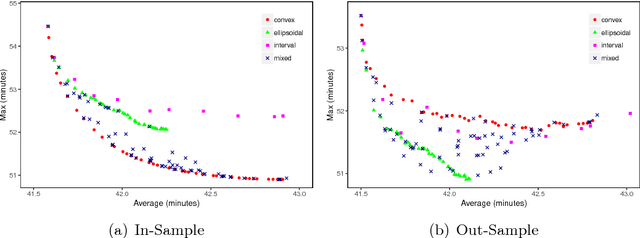
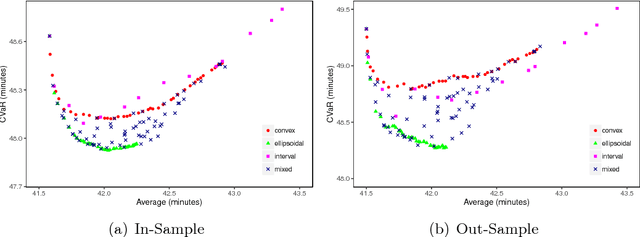
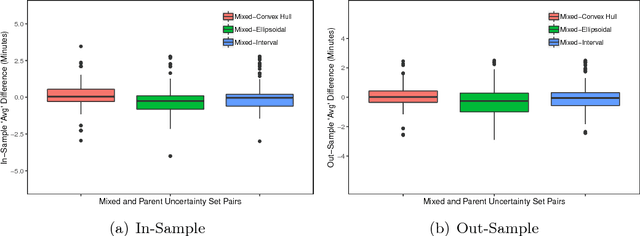
Abstract:In robust optimization, the uncertainty set is used to model all possible outcomes of uncertain parameters. In the classic setting, one assumes that this set is provided by the decision maker based on the data available to her. Only recently it has been recognized that the process of building useful uncertainty sets is in itself a challenging task that requires mathematical support. In this paper, we propose an approach to go beyond the classic setting, by assuming multiple uncertainty sets to be prepared, each with a weight showing the degree of belief that the set is a "true" model of uncertainty. We consider theoretical aspects of this approach and show that it is as easy to model as the classic setting. In an extensive computational study using a shortest path problem based on real-world data, we auto-tune uncertainty sets to the available data, and show that with regard to out-sample performance, the combination of multiple sets can give better results than each set on its own.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge