Quansong He
Enhancing Feature Fusion of U-like Networks with Dynamic Skip Connections
Sep 18, 2025Abstract:U-like networks have become fundamental frameworks in medical image segmentation through skip connections that bridge high-level semantics and low-level spatial details. Despite their success, conventional skip connections exhibit two key limitations: inter-feature constraints and intra-feature constraints. The inter-feature constraint refers to the static nature of feature fusion in traditional skip connections, where information is transmitted along fixed pathways regardless of feature content. The intra-feature constraint arises from the insufficient modeling of multi-scale feature interactions, thereby hindering the effective aggregation of global contextual information. To overcome these limitations, we propose a novel Dynamic Skip Connection (DSC) block that fundamentally enhances cross-layer connectivity through adaptive mechanisms. The DSC block integrates two complementary components. (1) Test-Time Training (TTT) module. This module addresses the inter-feature constraint by enabling dynamic adaptation of hidden representations during inference, facilitating content-aware feature refinement. (2) Dynamic Multi-Scale Kernel (DMSK) module. To mitigate the intra-feature constraint, this module adaptively selects kernel sizes based on global contextual cues, enhancing the network capacity for multi-scale feature integration. The DSC block is architecture-agnostic and can be seamlessly incorporated into existing U-like network structures. Extensive experiments demonstrate the plug-and-play effectiveness of the proposed DSC block across CNN-based, Transformer-based, hybrid CNN-Transformer, and Mamba-based U-like networks.
FuseUNet: A Multi-Scale Feature Fusion Method for U-like Networks
Jun 06, 2025Abstract:Medical image segmentation is a critical task in computer vision, with UNet serving as a milestone architecture. The typical component of UNet family is the skip connection, however, their skip connections face two significant limitations: (1) they lack effective interaction between features at different scales, and (2) they rely on simple concatenation or addition operations, which constrain efficient information integration. While recent improvements to UNet have focused on enhancing encoder and decoder capabilities, these limitations remain overlooked. To overcome these challenges, we propose a novel multi-scale feature fusion method that reimagines the UNet decoding process as solving an initial value problem (IVP), treating skip connections as discrete nodes. By leveraging principles from the linear multistep method, we propose an adaptive ordinary differential equation method to enable effective multi-scale feature fusion. Our approach is independent of the encoder and decoder architectures, making it adaptable to various U-Net-like networks. Experiments on ACDC, KiTS2023, MSD brain tumor, and ISIC2017/2018 skin lesion segmentation datasets demonstrate improved feature utilization, reduced network parameters, and maintained high performance. The code is available at https://github.com/nayutayuki/FuseUNet.
A Lightweight U-like Network Utilizing Neural Memory Ordinary Differential Equations for Slimming the Decoder
Dec 09, 2024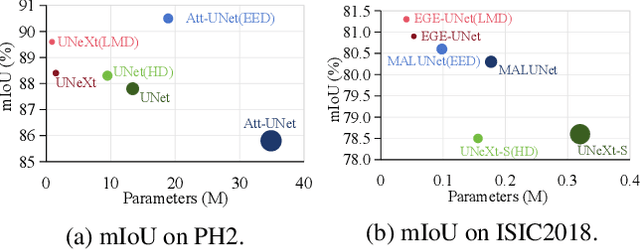
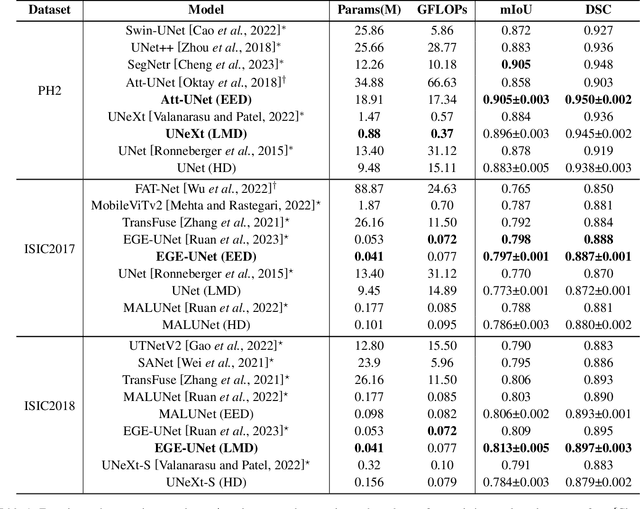
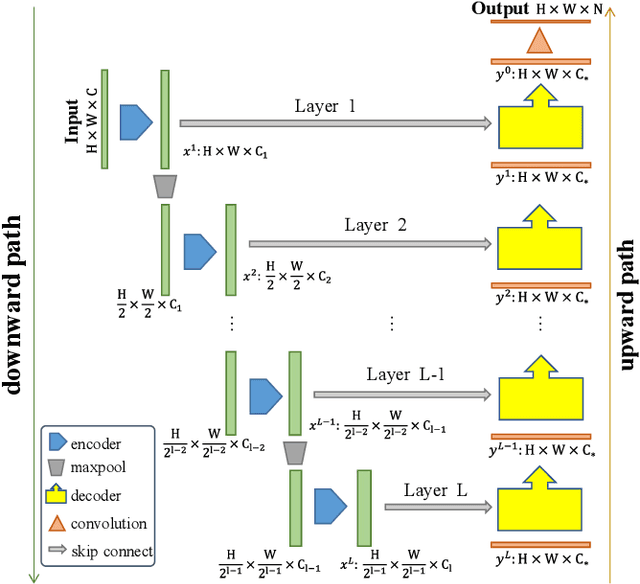
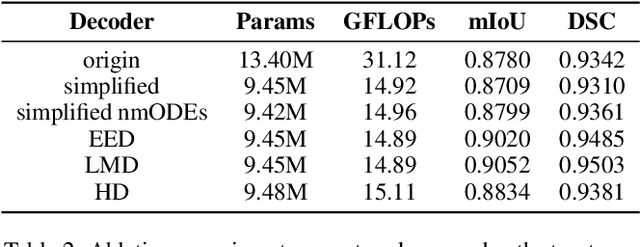
Abstract:In recent years, advanced U-like networks have demonstrated remarkable performance in medical image segmentation tasks. However, their drawbacks, including excessive parameters, high computational complexity, and slow inference speed, pose challenges for practical implementation in scenarios with limited computational resources. Existing lightweight U-like networks have alleviated some of these problems, but they often have pre-designed structures and consist of inseparable modules, limiting their application scenarios. In this paper, we propose three plug-and-play decoders by employing different discretization methods of the neural memory Ordinary Differential Equations (nmODEs). These decoders integrate features at various levels of abstraction by processing information from skip connections and performing numerical operations on upward path. Through experiments on the PH2, ISIC2017, and ISIC2018 datasets, we embed these decoders into different U-like networks, demonstrating their effectiveness in significantly reducing the number of parameters and FLOPs while maintaining performance. In summary, the proposed discretized nmODEs decoders are capable of reducing the number of parameters by about 20% ~ 50% and FLOPs by up to 74%, while possessing the potential to adapt to all U-like networks. Our code is available at https://github.com/nayutayuki/Lightweight-nmODE-Decoders-For-U-like-networks.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge