Qiao Zheng
Unsupervised shape and motion analysis of 3822 cardiac 4D MRIs of UK Biobank
Feb 15, 2019
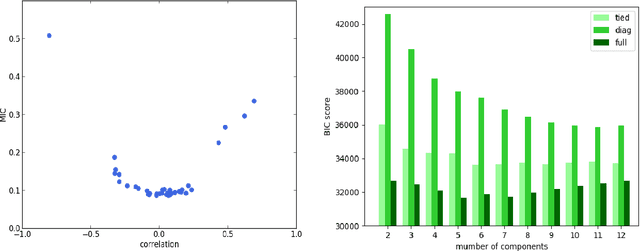
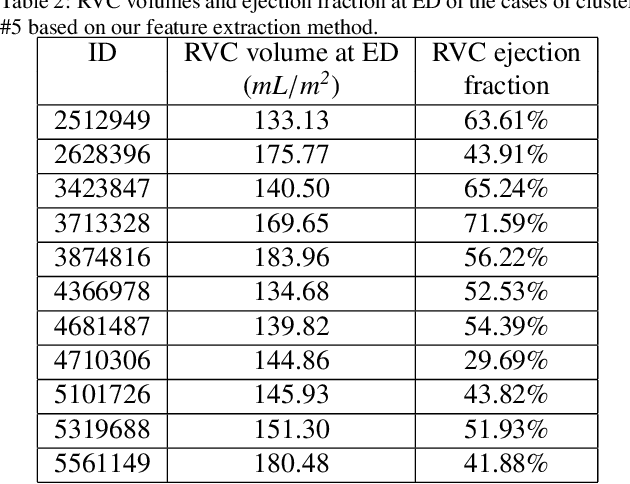
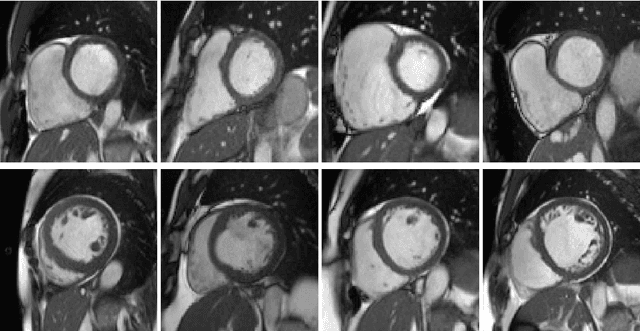
Abstract:We perform unsupervised analysis of image-derived shape and motion features extracted from 3822 cardiac 4D MRIs of the UK Biobank. First, with a feature extraction method previously published based on deep learning models, we extract from each case 9 feature values characterizing both the cardiac shape and motion. Second, a feature selection is performed to remove highly correlated feature pairs. Third, clustering is carried out using a Gaussian mixture model on the selected features. After analysis, we identify two small clusters which probably correspond to two pathological categories. Further confirmation using a trained classification model and dimensionality reduction tools is carried out to support this discovery. Moreover, we examine the differences between the other large clusters and compare our measures with the ground-truth.
Explainable cardiac pathology classification on cine MRI with motion characterization by semi-supervised learning of apparent flow
Nov 08, 2018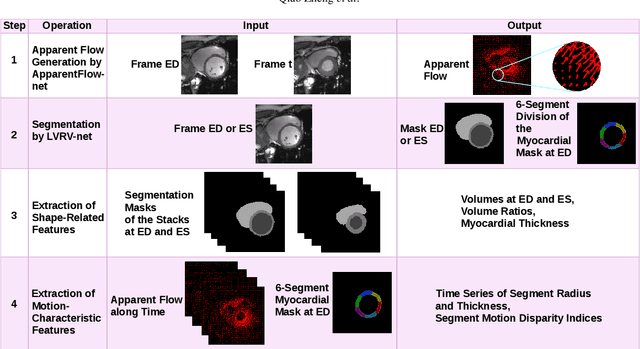

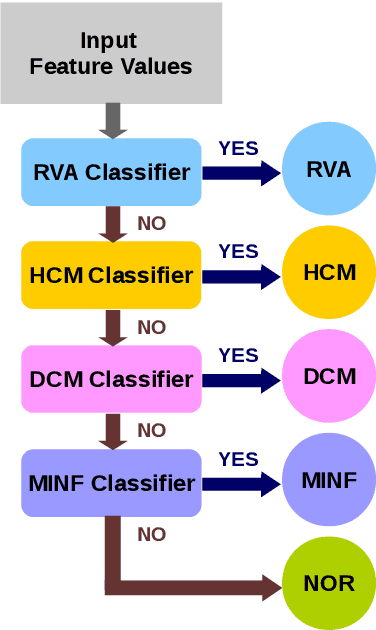

Abstract:We propose a method to classify cardiac pathology based on a novel approach to extract image derived features to characterize the shape and motion of the heart. An original semi-supervised learning procedure, which makes efficient use of a large amount of non-segmented images and a small amount of images segmented manually by experts, is developed to generate pixel-wise apparent flow between two time points of a 2D+t cine MRI image sequence. Combining the apparent flow maps and cardiac segmentation masks, we obtain a local apparent flow corresponding to the 2D motion of myocardium and ventricular cavities. This leads to the generation of time series of the radius and thickness of myocardial segments to represent cardiac motion. These time series of motion features are reliable and explainable characteristics of pathological cardiac motion. Furthermore, they are combined with shape-related features to classify cardiac pathologies. Using only nine feature values as input, we propose an explainable, simple and flexible model for pathology classification. On ACDC training set and testing set, the model achieves 95% and 94% respectively as classification accuracy. Its performance is hence comparable to that of the state-of-the-art. Comparison with various other models is performed to outline some advantages of our model.
3D Consistent & Robust Segmentation of Cardiac Images by Deep Learning with Spatial Propagation
Apr 25, 2018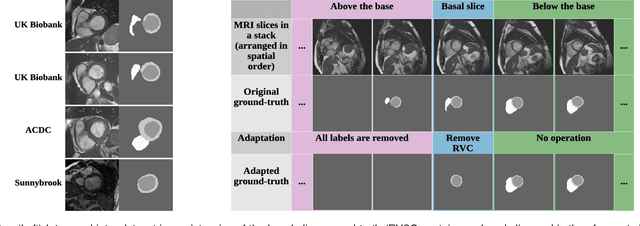
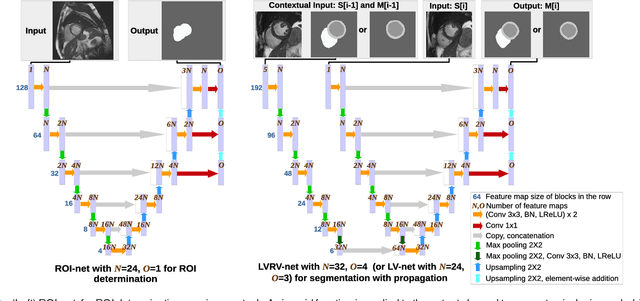
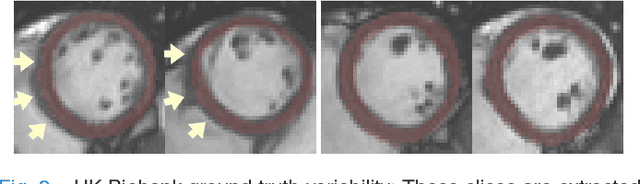
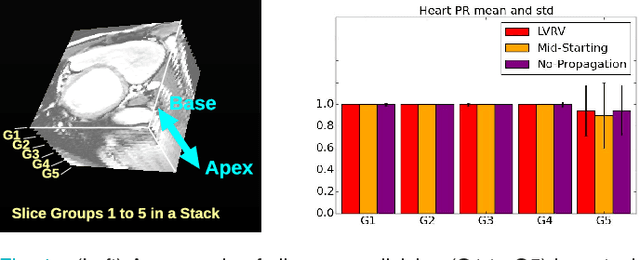
Abstract:We propose a method based on deep learning to perform cardiac segmentation on short axis MRI image stacks iteratively from the top slice (around the base) to the bottom slice (around the apex). At each iteration, a novel variant of U-net is applied to propagate the segmentation of a slice to the adjacent slice below it. In other words, the prediction of a segmentation of a slice is dependent upon the already existing segmentation of an adjacent slice. 3D-consistency is hence explicitly enforced. The method is trained on a large database of 3078 cases from UK Biobank. It is then tested on 756 different cases from UK Biobank and three other state-of-the-art cohorts (ACDC with 100 cases, Sunnybrook with 30 cases, RVSC with 16 cases). Results comparable or even better than the state-of-the-art in terms of distance measures are achieved. They also emphasize the assets of our method, namely enhanced spatial consistency (currently neither considered nor achieved by the state-of-the-art), and the generalization ability to unseen cases even from other databases.
3D Consistent Biventricular Myocardial Segmentation Using Deep Learning for Mesh Generation
Mar 29, 2018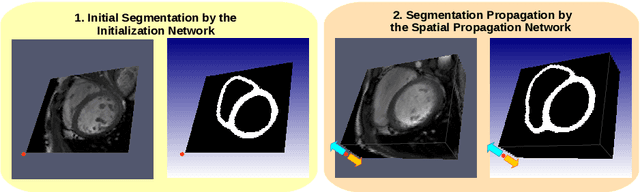
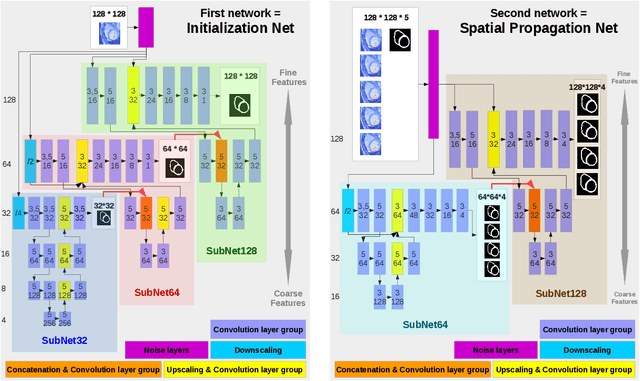
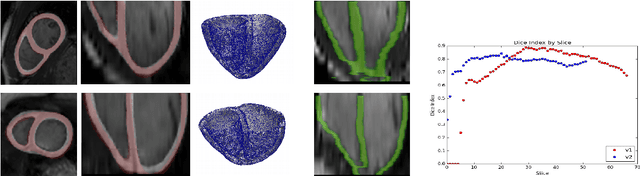
Abstract:We present a novel automated method to segment the myocardium of both left and right ventricles in MRI volumes. The segmentation is consistent in 3D across the slices such that it can be directly used for mesh generation. Two specific neural networks with multi-scale coarse-to-fine prediction structure are proposed to cope with the small training dataset and trained using an original loss function. The former segments a slice in the middle of the volume. Then the latter iteratively propagates the slice segmentations towards the base and the apex, in a spatially consistent way. We perform 5-fold cross-validation on the 15 cases from STACOM to validate the method. For training, we use real cases and their synthetic variants generated by combining motion simulation and image synthesis. Accurate and consistent testing results are obtained.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge