Olli Gröhn
RatLesNetv2: A Fully Convolutional Network for Rodent Brain Lesion Segmentation
Jan 24, 2020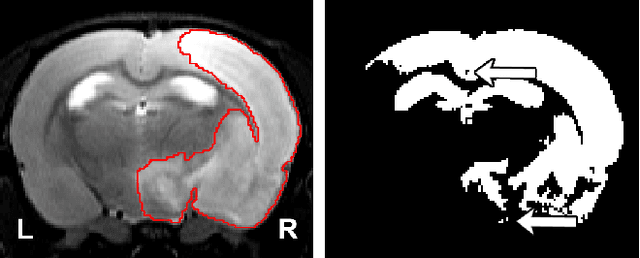
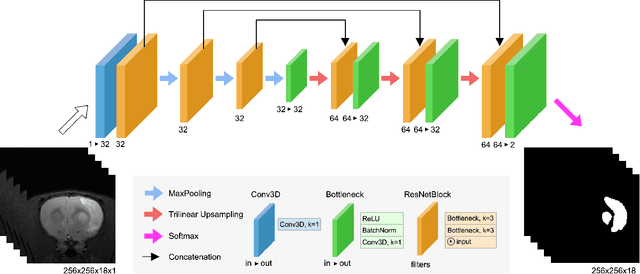

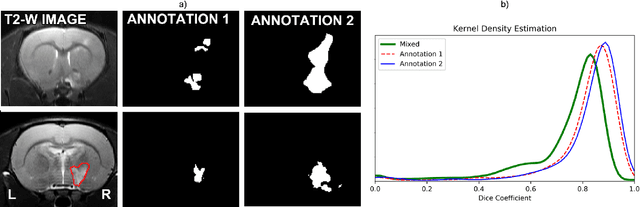
Abstract:Segmentation of rodent brain lesions on magnetic resonance images (MRIs) is a time-consuming task with high inter- and intra-operator variability due to its subjective nature. We present a three-dimensional fully convolutional neural network (ConvNet) named RatLesNetv2 for segmenting rodent brain lesions. We compare its performance with other ConvNets on an unusually large and heterogeneous data set composed by 916 T2-weighted rat brain scans at nine different lesion stages. RatLesNetv2 obtained similar to higher Dice coefficients than the other ConvNets and it produced much more realistic and compact segmentations with notably less holes and lower Hausdorff distance. RatLesNetv2-derived segmentations also exceeded inter-rater agreement Dice coefficients. Additionally, we show that training on disparate ground truths leads to significantly different segmentations, and we study RatLesNetv2 generalization capability when optimizing for training sets of different sizes. RatLesNetv2 is publicly available at https://github.com/jmlipman/RatLesNetv2.
Automatic Rodent Brain MRI Lesion Segmentation with Fully Convolutional Networks
Aug 23, 2019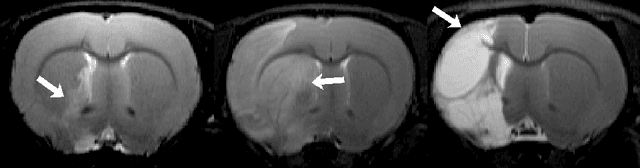
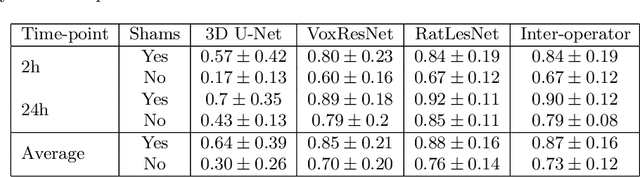
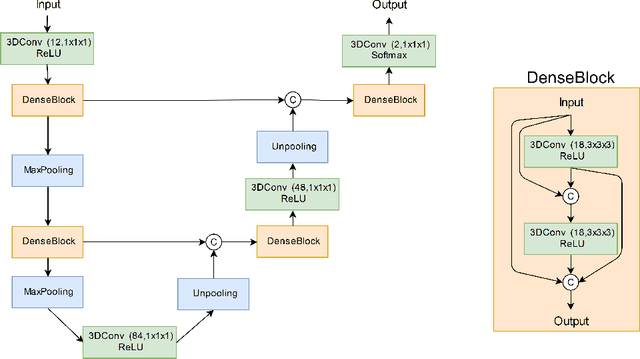
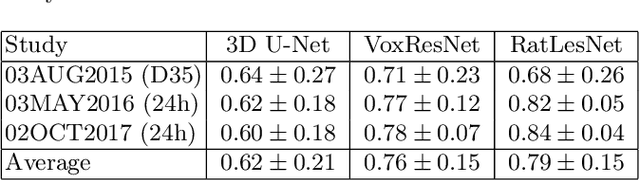
Abstract:Manual segmentation of rodent brain lesions from magnetic resonance images (MRIs) is an arduous, time-consuming and subjective task that is highly important in pre-clinical research. Several automatic methods have been developed for different human brain MRI segmentation, but little research has targeted automatic rodent lesion segmentation. The existing tools for performing automatic lesion segmentation in rodents are constrained by strict assumptions about the data. Deep learning has been successfully used for medical image segmentation. However, there has not been any deep learning approach specifically designed for tackling rodent brain lesion segmentation. In this work, we propose a novel Fully Convolutional Network (FCN), RatLesNet, for the aforementioned task. Our dataset consists of 131 T2-weighted rat brain scans from 4 different studies in which ischemic stroke was induced by transient middle cerebral artery occlusion. We compare our method with two other 3D FCNs originally developed for anatomical segmentation (VoxResNet and 3D-U-Net) with 5-fold cross-validation on a single study and a generalization test, where the training was done on a single study and testing on three remaining studies. The labels generated by our method were quantitatively and qualitatively better than the predictions of the compared methods. The average Dice coefficient achieved in the 5-fold cross-validation experiment with the proposed approach was 0.88, between 3.7% and 38% higher than the compared architectures. The presented architecture also outperformed the other FCNs at generalizing on different studies, achieving the average Dice coefficient of 0.79.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge