Nipuna Senanayake
A Survey on Graph Neural Networks for Fraud Detection in Ride Hailing Platforms
Dec 29, 2025Abstract:This study investigates fraud detection in ride hailing platforms through Graph Neural Networks (GNNs),focusing on the effectiveness of various models. By analyzing prevalent fraudulent activities, the research highlights and compares the existing work related to fraud detection which can be useful when addressing fraudulent incidents within the online ride hailing platforms. Also, the paper highlights addressing class imbalance and fraudulent camouflage. It also outlines a structured overview of GNN architectures and methodologies applied to anomaly detection, identifying significant methodological progress and gaps. The paper calls for further exploration into real-world applicability and technical improvements to enhance fraud detection strategies in the rapidly evolving ride-hailing industry.
* 12 pages, 8 figures, 2 tables. Presented at the 2024 7th International Conference on Artificial Intelligence and Big Data (ICAIBD)
Advancements in Natural Language Processing for Automatic Text Summarization
Feb 27, 2025



Abstract:The substantial growth of textual content in diverse domains and platforms has led to a considerable need for Automatic Text Summarization (ATS) techniques that aid in the process of text analysis. The effectiveness of text summarization models has been significantly enhanced in a variety of technical domains because of advancements in Natural Language Processing (NLP) and Deep Learning (DL). Despite this, the process of summarizing textual information continues to be significantly constrained by the intricate writing styles of a variety of texts, which involve a range of technical complexities. Text summarization techniques can be broadly categorized into two main types: abstractive summarization and extractive summarization. Extractive summarization involves directly extracting sentences, phrases, or segments of text from the content without making any changes. On the other hand, abstractive summarization is achieved by reconstructing the sentences, phrases, or segments from the original text using linguistic analysis. Through this study, a linguistically diverse categorizations of text summarization approaches have been addressed in a constructive manner. In this paper, the authors explored existing hybrid techniques that have employed both extractive and abstractive methodologies. In addition, the pros and cons of various approaches discussed in the literature are also investigated. Furthermore, the authors conducted a comparative analysis on different techniques and matrices to evaluate the generated summaries using language generation models. This survey endeavors to provide a comprehensive overview of ATS by presenting the progression of language processing regarding this task through a breakdown of diverse systems and architectures accompanied by technical and mathematical explanations of their operations.
* 11 pages, 9 figures, ICCS 2024
Covid-19 Detection from Chest X-ray and Patient Metadata using Graph Convolutional Neural Networks
May 21, 2021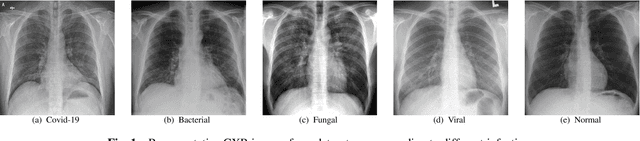
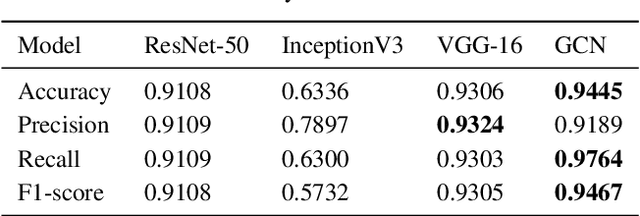
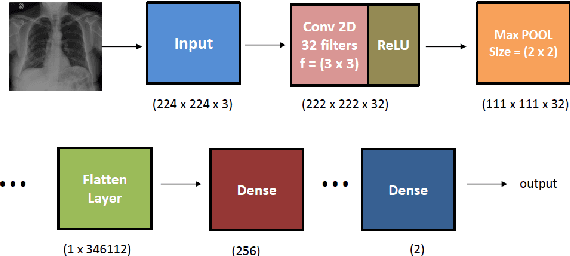
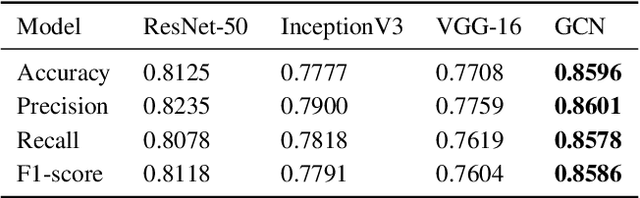
Abstract:The novel corona virus (Covid-19) has introduced significant challenges due to its rapid spreading nature through respiratory transmission. As a result, there is a huge demand for Artificial Intelligence (AI) based quick disease diagnosis methods as an alternative to high demand tests such as Polymerase Chain Reaction (PCR). Chest X-ray (CXR) Image analysis is such cost-effective radiography technique due to resource availability and quick screening. But, a sufficient and systematic data collection that is required by complex deep leaning (DL) models is more difficult and hence there are recent efforts that utilize transfer learning to address this issue. Still these transfer learnt models suffer from lack of generalization and increased bias to the training dataset resulting poor performance for unseen data. Limited correlation of the transferred features from the pre-trained model to a specific medical imaging domain like X-ray and overfitting on fewer data can be reasons for this circumstance. In this work, we propose a novel Graph Convolution Neural Network (GCN) that is capable of identifying bio-markers of Covid-19 pneumonia from CXR images and meta information about patients. The proposed method exploits important relational knowledge between data instances and their features using graph representation and applies convolution to learn the graph data which is not possible with conventional convolution on Euclidean domain. The results of extensive experiments of proposed model on binary (Covid vs normal) and three class (Covid, normal, other pneumonia) classification problems outperform different benchmark transfer learnt models, hence overcoming the aforementioned drawbacks.
Graph Convolution Networks Using Message Passing and Multi-Source Similarity Features for Predicting circRNA-Disease Association
Sep 15, 2020

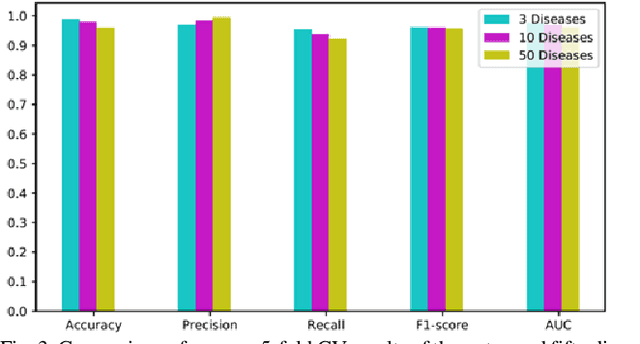
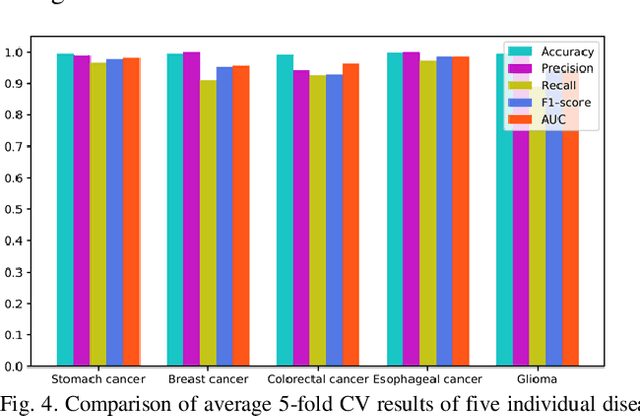
Abstract:Graphs can be used to effectively represent complex data structures. Learning these irregular data in graphs is challenging and still suffers from shallow learning. Applying deep learning on graphs has recently showed good performance in many applications in social analysis, bioinformatics etc. A message passing graph convolution network is such a powerful method which has expressive power to learn graph structures. Meanwhile, circRNA is a type of non-coding RNA which plays a critical role in human diseases. Identifying the associations between circRNAs and diseases is important to diagnosis and treatment of complex diseases. However, there are limited number of known associations between them and conducting biological experiments to identify new associations is time consuming and expensive. As a result, there is a need of building efficient and feasible computation methods to predict potential circRNA-disease associations. In this paper, we propose a novel graph convolution network framework to learn features from a graph built with multi-source similarity information to predict circRNA-disease associations. First we use multi-source information of circRNA similarity, disease and circRNA Gaussian Interaction Profile (GIP) kernel similarity to extract the features using first graph convolution. Then we predict disease associations for each circRNA with second graph convolution. Proposed framework with five-fold cross validation on various experiments shows promising results in predicting circRNA-disease association and outperforms other existing methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge