Nathalie Willems
Semi-Supervised Segmentation via Embedding Matching
Jul 05, 2024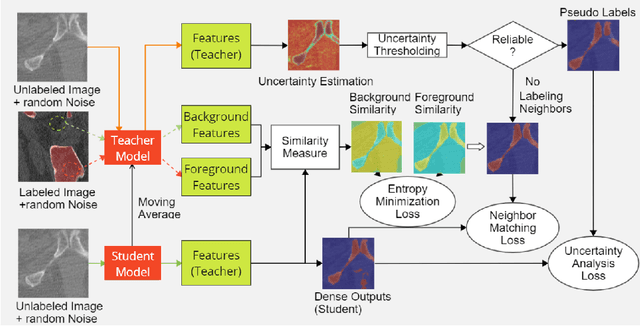
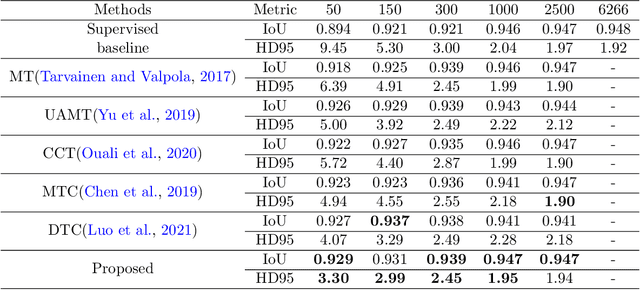
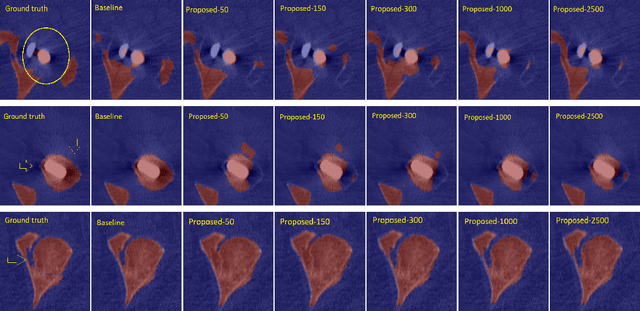

Abstract:Deep convolutional neural networks are widely used in medical image segmentation but require many labeled images for training. Annotating three-dimensional medical images is a time-consuming and costly process. To overcome this limitation, we propose a novel semi-supervised segmentation method that leverages mostly unlabeled images and a small set of labeled images in training. Our approach involves assessing prediction uncertainty to identify reliable predictions on unlabeled voxels from the teacher model. These voxels serve as pseudo-labels for training the student model. In voxels where the teacher model produces unreliable predictions, pseudo-labeling is carried out based on voxel-wise embedding correspondence using reference voxels from labeled images. We applied this method to automate hip bone segmentation in CT images, achieving notable results with just 4 CT scans. The proposed approach yielded a Hausdorff distance with 95th percentile (HD95) of 3.30 and IoU of 0.929, surpassing existing methods achieving HD95 (4.07) and IoU (0.927) at their best.
SAM Fewshot Finetuning for Anatomical Segmentation in Medical Images
Jul 05, 2024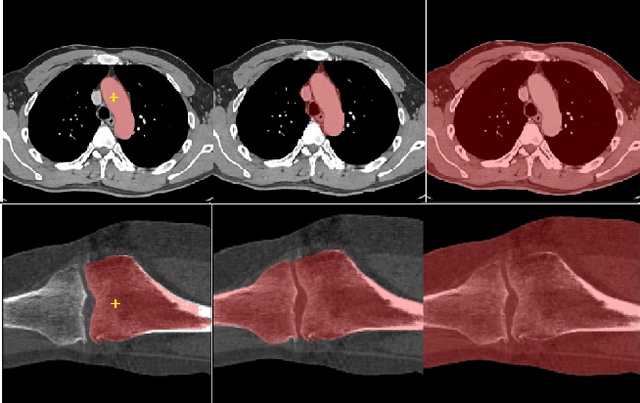
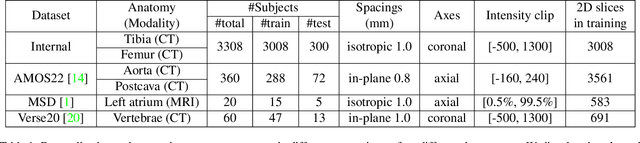
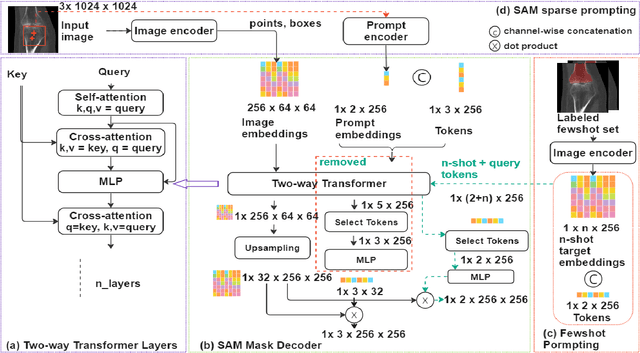
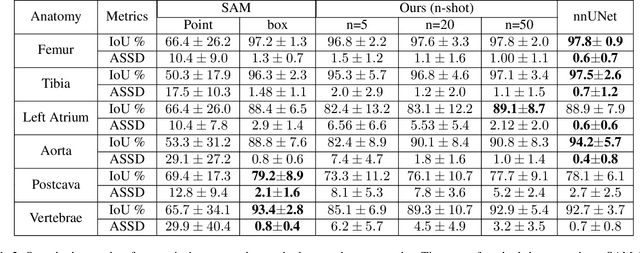
Abstract:We propose a straightforward yet highly effective few-shot fine-tuning strategy for adapting the Segment Anything (SAM) to anatomical segmentation tasks in medical images. Our novel approach revolves around reformulating the mask decoder within SAM, leveraging few-shot embeddings derived from a limited set of labeled images (few-shot collection) as prompts for querying anatomical objects captured in image embeddings. This innovative reformulation greatly reduces the need for time-consuming online user interactions for labeling volumetric images, such as exhaustively marking points and bounding boxes to provide prompts slice by slice. With our method, users can manually segment a few 2D slices offline, and the embeddings of these annotated image regions serve as effective prompts for online segmentation tasks. Our method prioritizes the efficiency of the fine-tuning process by exclusively training the mask decoder through caching mechanisms while keeping the image encoder frozen. Importantly, this approach is not limited to volumetric medical images, but can generically be applied to any 2D/3D segmentation task. To thoroughly evaluate our method, we conducted extensive validation on four datasets, covering six anatomical segmentation tasks across two modalities. Furthermore, we conducted a comparative analysis of different prompting options within SAM and the fully-supervised nnU-Net. The results demonstrate the superior performance of our method compared to SAM employing only point prompts (approximately 50% improvement in IoU) and performs on-par with fully supervised methods whilst reducing the requirement of labeled data by at least an order of magnitude.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge