Núria Queralt-Rosinach
Synthetic data: How could it be used for infectious disease research?
Jul 03, 2024Abstract:Over the last three to five years, it has become possible to generate machine learning synthetic data for healthcare-related uses. However, concerns have been raised about potential negative factors associated with the possibilities of artificial dataset generation. These include the potential misuse of generative artificial intelligence (AI) in fields such as cybercrime, the use of deepfakes and fake news to deceive or manipulate, and displacement of human jobs across various market sectors. Here, we consider both current and future positive advances and possibilities with synthetic datasets. Synthetic data offers significant benefits, particularly in data privacy, research, in balancing datasets and reducing bias in machine learning models. Generative AI is an artificial intelligence genre capable of creating text, images, video or other data using generative models. The recent explosion of interest in GenAI was heralded by the invention and speedy move to use of large language models (LLM). These computational models are able to achieve general-purpose language generation and other natural language processing tasks and are based on transformer architectures, which made an evolutionary leap from previous neural network architectures. Fuelled by the advent of improved GenAI techniques and wide scale usage, this is surely the time to consider how synthetic data can be used to advance infectious disease research. In this commentary we aim to create an overview of the current and future position of synthetic data in infectious disease research.
Neuro-symbolic representation learning on biological knowledge graphs
Dec 13, 2016
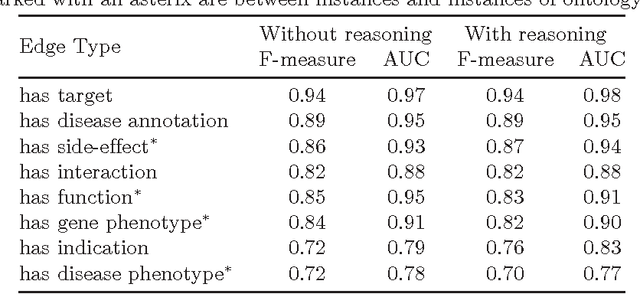
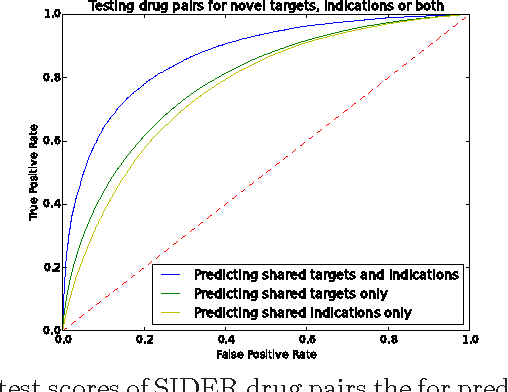
Abstract:Motivation: Biological data and knowledge bases increasingly rely on Semantic Web technologies and the use of knowledge graphs for data integration, retrieval and federated queries. In the past years, feature learning methods that are applicable to graph-structured data are becoming available, but have not yet widely been applied and evaluated on structured biological knowledge. Results: We develop a novel method for feature learning on biological knowledge graphs. Our method combines symbolic methods, in particular knowledge representation using symbolic logic and automated reasoning, with neural networks to generate embeddings of nodes that encode for related information within knowledge graphs. Through the use of symbolic logic, these embeddings contain both explicit and implicit information. We apply these embeddings to the prediction of edges in the knowledge graph representing problems of function prediction, finding candidate genes of diseases, protein-protein interactions, or drug target relations, and demonstrate performance that matches and sometimes outperforms traditional approaches based on manually crafted features. Our method can be applied to any biological knowledge graph, and will thereby open up the increasing amount of Semantic Web based knowledge bases in biology to use in machine learning and data analytics. Availability and Implementation: https://github.com/bio-ontology-research-group/walking-rdf-and-owl Contact: robert.hoehndorf@kaust.edu.sa
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge