Michela Tosetti
MRI2Qmap: multi-parametric quantitative mapping with MRI-driven denoising priors
Mar 11, 2026Abstract:Magnetic Resonance Fingerprinting (MRF) and other highly accelerated transient-state parameter mapping techniques enable simultaneous quantification of multiple tissue properties, but often suffer from aliasing artifacts due to compressed sampling. Incorporating spatial image priors can mitigate these artifacts, and deep learning has shown strong potential when large training datasets are available. However, extending this paradigm to MRF-type sequences remains challenging due to the scarcity of quantitative imaging data for training. Can this limitation be overcome by leveraging sources of training data from clinically-routine weighted MRI images? To this end, we introduce MRI2Qmap, a plug-and-play quantitative reconstruction framework that integrates the physical acquisition model with priors learned from deep denoising autoencoders pretrained on large multimodal weighted-MRI datasets. MRI2Qmap demonstrates that spatial-domain structural priors learned from independently acquired datasets of routine weighted-MRI images can be effectively used for quantitative MRI reconstruction. The proposed method is validated on highly accelerated 3D whole-brain MRF data from both in-vivo and simulated acquisitions, achieving competitive or superior performance relative to existing baselines without requiring ground-truth quantitative imaging data for training. By decoupling quantitative reconstruction from the need for ground-truth MRF training data, this framework points toward a scalable paradigm for quantitative MRI that can capitalize on the large and growing repositories of routine clinical MRI.
StoDIP: Efficient 3D MRF image reconstruction with deep image priors and stochastic iterations
Aug 05, 2024


Abstract:Magnetic Resonance Fingerprinting (MRF) is a time-efficient approach to quantitative MRI for multiparametric tissue mapping. The reconstruction of quantitative maps requires tailored algorithms for removing aliasing artefacts from the compressed sampled MRF acquisitions. Within approaches found in the literature, many focus solely on two-dimensional (2D) image reconstruction, neglecting the extension to volumetric (3D) scans despite their higher relevance and clinical value. A reason for this is that transitioning to 3D imaging without appropriate mitigations presents significant challenges, including increased computational cost and storage requirements, and the need for large amount of ground-truth (artefact-free) data for training. To address these issues, we introduce StoDIP, a new algorithm that extends the ground-truth-free Deep Image Prior (DIP) reconstruction to 3D MRF imaging. StoDIP employs memory-efficient stochastic updates across the multicoil MRF data, a carefully selected neural network architecture, as well as faster nonuniform FFT (NUFFT) transformations. This enables a faster convergence compared against a conventional DIP implementation without these features. Tested on a dataset of whole-brain scans from healthy volunteers, StoDIP demonstrated superior performance over the ground-truth-free reconstruction baselines, both quantitatively and qualitatively.
Deep Image Priors for Magnetic Resonance Fingerprinting with pretrained Bloch-consistent denoising autoencoders
Jul 29, 2024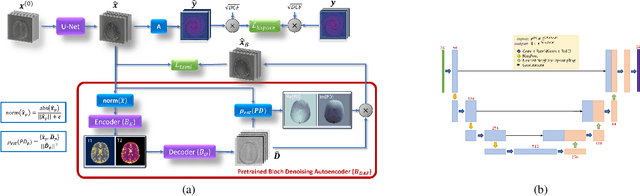
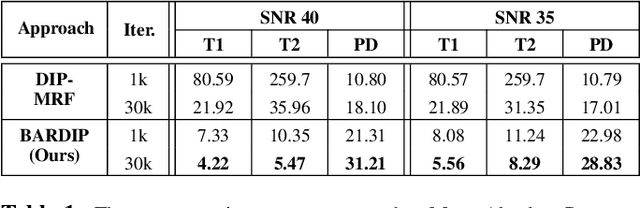

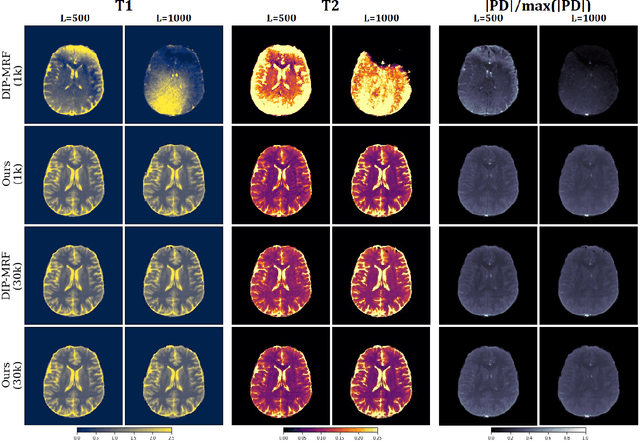
Abstract:The estimation of multi-parametric quantitative maps from Magnetic Resonance Fingerprinting (MRF) compressed sampled acquisitions, albeit successful, remains a challenge due to the high underspampling rate and artifacts naturally occuring during image reconstruction. Whilst state-of-the-art DL methods can successfully address the task, to fully exploit their capabilities they often require training on a paired dataset, in an area where ground truth is seldom available. In this work, we propose a method that combines a deep image prior (DIP) module that, without ground truth and in conjunction with a Bloch consistency enforcing autoencoder, can tackle the problem, resulting in a method faster and of equivalent or better accuracy than DIP-MRF.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge