Mengyu Sun
Trust the Unreliability: Inward Backward Dynamic Unreliability Driven Coreset Selection for Medical Image Classification
Mar 18, 2026Abstract:Efficiently managing and utilizing large-scale medical imaging datasets with limited resources presents significant challenges. While coreset selection helps reduce computational costs, its effectiveness in medical data remains limited due to inherent complexity, such as large intra-class variation and high inter-class similarity. To address this, we revisit the training process and observe that neural networks consistently produce stable confidence predictions and better remember samples near class centers in training. However, concentrating on these samples may complicate the modeling of decision boundaries. Hence, we argue that the more unreliable samples are, in fact, the more informative in helping build the decision boundary. Based on this, we propose the Dynamic Unreliability-Driven Coreset Selection(DUCS) strategy. Specifically, we introduce an inward-backward unreliability assessment perspective: 1) Inward Self-Awareness: The model introspects its behavior by analyzing the evolution of confidence during training, thereby quantifying uncertainty of each sample. 2) Backward Memory Tracking: The model reflects on its training tracking by tracking the frequency of forgetting samples, thus evaluating its retention ability for each sample. Next, we select unreliable samples that exhibit substantial confidence fluctuations and are repeatedly forgotten during training. This selection process ensures that the chosen samples are near the decision boundary, thereby aiding the model in refining the boundary. Extensive experiments on public medical datasets demonstrate our superior performance compared to state-of-the-art(SOTA) methods, particularly at high compression rates.
Code Fingerprints: Disentangled Attribution of LLM-Generated Code
Mar 04, 2026Abstract:The rapid adoption of Large Language Models (LLMs) has transformed modern software development by enabling automated code generation at scale. While these systems improve productivity, they introduce new challenges for software governance, accountability, and compliance. Existing research primarily focuses on distinguishing machine-generated code from human-written code; however, many practical scenarios--such as vulnerability triage, incident investigation, and licensing audits--require identifying which LLM produced a given code snippet. In this paper, we study the problem of model-level code attribution, which aims to determine the source LLM responsible for generated code. Although attribution is challenging, differences in training data, architectures, alignment strategies, and decoding mechanisms introduce model-dependent stylistic and structural variations that serve as generative fingerprints. Leveraging this observation, we propose the Disentangled Code Attribution Network (DCAN), which separates Source-Agnostic semantic information from Source-Specific stylistic representations. Through a contrastive learning objective, DCAN isolates discriminative model-dependent signals while preserving task semantics, enabling multi-class attribution across models and programming languages. To support systematic evaluation, we construct the first large-scale benchmark dataset comprising code generated by four widely used LLMs (DeepSeek, Claude, Qwen, and ChatGPT) across four programming languages (Python, Java, C, and Go). Experimental results demonstrate that DCAN achieves reliable attribution performance across diverse settings, highlighting the feasibility of model-level provenance analysis in software engineering contexts. The dataset and implementation are publicly available at https://github.com/mtt500/DCAN.
LURE: Latent Space Unblocking for Multi-Concept Reawakening in Diffusion Models
Jan 20, 2026Abstract:Concept erasure aims to suppress sensitive content in diffusion models, but recent studies show that erased concepts can still be reawakened, revealing vulnerabilities in erasure methods. Existing reawakening methods mainly rely on prompt-level optimization to manipulate sampling trajectories, neglecting other generative factors, which limits a comprehensive understanding of the underlying dynamics. In this paper, we model the generation process as an implicit function to enable a comprehensive theoretical analysis of multiple factors, including text conditions, model parameters, and latent states. We theoretically show that perturbing each factor can reawaken erased concepts. Building on this insight, we propose a novel concept reawakening method: Latent space Unblocking for concept REawakening (LURE), which reawakens erased concepts by reconstructing the latent space and guiding the sampling trajectory. Specifically, our semantic re-binding mechanism reconstructs the latent space by aligning denoising predictions with target distributions to reestablish severed text-visual associations. However, in multi-concept scenarios, naive reconstruction can cause gradient conflicts and feature entanglement. To address this, we introduce Gradient Field Orthogonalization, which enforces feature orthogonality to prevent mutual interference. Additionally, our Latent Semantic Identification-Guided Sampling (LSIS) ensures stability of the reawakening process via posterior density verification. Extensive experiments demonstrate that LURE enables simultaneous, high-fidelity reawakening of multiple erased concepts across diverse erasure tasks and methods.
Low-Level Dataset Distillation for Medical Image Enhancement
Nov 17, 2025Abstract:Medical image enhancement is clinically valuable, but existing methods require large-scale datasets to learn complex pixel-level mappings. However, the substantial training and storage costs associated with these datasets hinder their practical deployment. While dataset distillation (DD) can alleviate these burdens, existing methods mainly target high-level tasks, where multiple samples share the same label. This many-to-one mapping allows distilled data to capture shared semantics and achieve information compression. In contrast, low-level tasks involve a many-to-many mapping that requires pixel-level fidelity, making low-level DD an underdetermined problem, as a small distilled dataset cannot fully constrain the dense pixel-level mappings. To address this, we propose the first low-level DD method for medical image enhancement. We first leverage anatomical similarities across patients to construct the shared anatomical prior based on a representative patient, which serves as the initialization for the distilled data of different patients. This prior is then personalized for each patient using a Structure-Preserving Personalized Generation (SPG) module, which integrates patient-specific anatomical information into the distilled dataset while preserving pixel-level fidelity. For different low-level tasks, the distilled data is used to construct task-specific high- and low-quality training pairs. Patient-specific knowledge is injected into the distilled data by aligning the gradients computed from networks trained on the distilled pairs with those from the corresponding patient's raw data. Notably, downstream users cannot access raw patient data. Instead, only a distilled dataset containing abstract training information is shared, which excludes patient-specific details and thus preserves privacy.
A CT Image Denoising Method Based on Projection Domain Feature
Dec 09, 2024
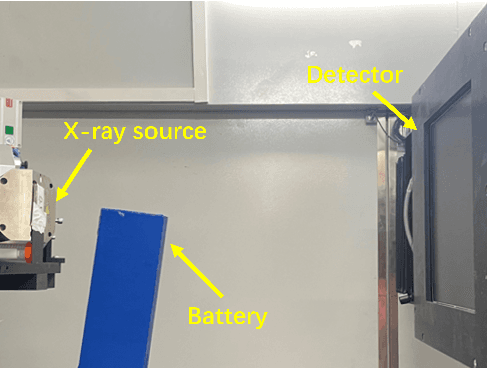



Abstract:In order to improve image quality of projection in industrial applications, generally, a standard method is to increase the current or exposure time, which might cause overexposure of detector units in areas of thin objects or backgrounds. Increasing the projection sampling is a better method to address the issue, but it also leads to significant noise in the reconstructed image. This paper proposed a projection domain denoising algorithm based on the features of the projection domain for this case. This algorithm utilized the similarity of projections of neighboring veiws to reduce image noise quickly and effectively. The availability of the algorithm proposed in this work has been conducted by numerical simulation and practical data experiments.
Plaintext-Free Deep Learning for Privacy-Preserving Medical Image Analysis via Frequency Information Embedding
Mar 25, 2024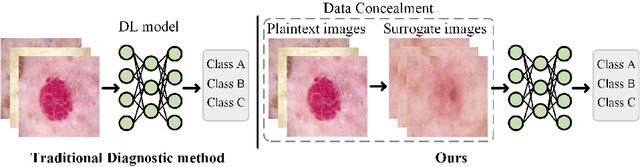
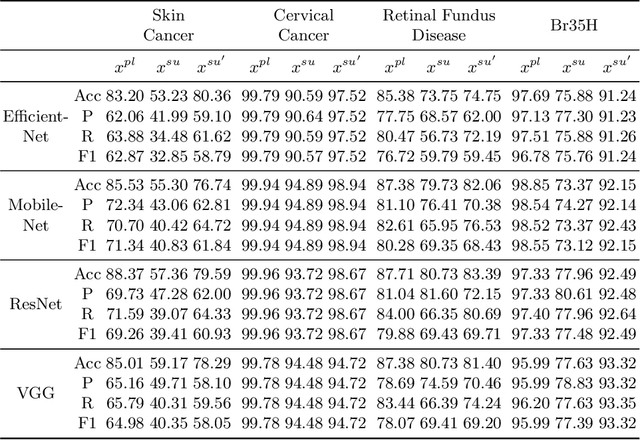
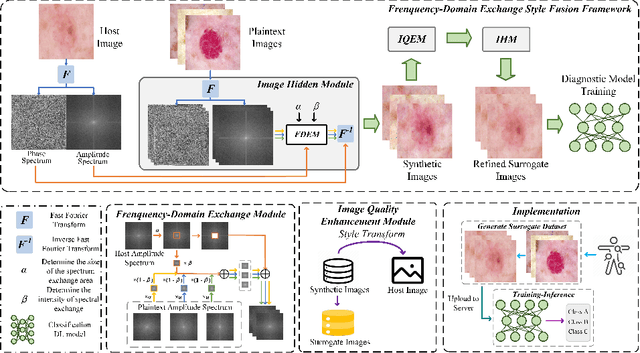
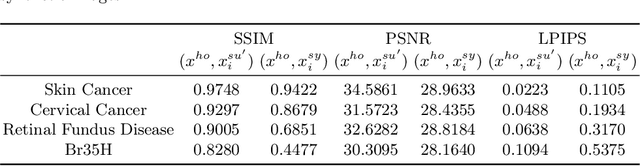
Abstract:In the fast-evolving field of medical image analysis, Deep Learning (DL)-based methods have achieved tremendous success. However, these methods require plaintext data for training and inference stages, raising privacy concerns, especially in the sensitive area of medical data. To tackle these concerns, this paper proposes a novel framework that uses surrogate images for analysis, eliminating the need for plaintext images. This approach is called Frequency-domain Exchange Style Fusion (FESF). The framework includes two main components: Image Hidden Module (IHM) and Image Quality Enhancement Module~(IQEM). The~IHM performs in the frequency domain, blending the features of plaintext medical images into host medical images, and then combines this with IQEM to improve and create surrogate images effectively. During the diagnostic model training process, only surrogate images are used, enabling anonymous analysis without any plaintext data during both training and inference stages. Extensive evaluations demonstrate that our framework effectively preserves the privacy of medical images and maintains diagnostic accuracy of DL models at a relatively high level, proving its effectiveness across various datasets and DL-based models.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge