May D Wang
KindSleep: Knowledge-Informed Diagnosis of Obstructive Sleep Apnea from Oximetry
Mar 05, 2026Abstract:Obstructive sleep apnea (OSA) is a sleep disorder that affects nearly one billion people globally and significantly elevates cardiovascular risk. Traditional diagnosis through polysomnography is resource-intensive and limits widespread access, creating a critical need for accurate and efficient alternatives. In this paper, we introduce KindSleep, a deep learning framework that integrates clinical knowledge with single-channel patient-specific oximetry signals and clinical data for precise OSA diagnosis. KindSleep first learns to identify clinically interpretable concepts, such as desaturation indices and respiratory disturbance events, directly from raw oximetry signals. It then fuses these AI-derived concepts with multimodal clinical data to estimate the Apnea-Hypopnea Index (AHI). We evaluate KindSleep on three large, independent datasets from the National Sleep Research Resource (SHHS, CFS, MrOS; total n = 9,815). KindSleep demonstrates excellent performance in estimating AHI scores (R2 = 0.917, ICC = 0.957) and consistently outperforms existing approaches in classifying OSA severity, achieving weighted F1-scores from 0.827 to 0.941 across diverse populations. By grounding its predictions in a layer of clinically meaningful concepts, KindSleep provides a more transparent and trustworthy diagnostic tool for sleep medicine practices.
Autonomous Soft Tissue Retraction Using Demonstration-Guided Reinforcement Learning
Sep 02, 2023Abstract:In the context of surgery, robots can provide substantial assistance by performing small, repetitive tasks such as suturing, needle exchange, and tissue retraction, thereby enabling surgeons to concentrate on more complex aspects of the procedure. However, existing surgical task learning mainly pertains to rigid body interactions, whereas the advancement towards more sophisticated surgical robots necessitates the manipulation of soft bodies. Previous work focused on tissue phantoms for soft tissue task learning, which can be expensive and can be an entry barrier to research. Simulation environments present a safe and efficient way to learn surgical tasks before their application to actual tissue. In this study, we create a Robot Operating System (ROS)-compatible physics simulation environment with support for both rigid and soft body interactions within surgical tasks. Furthermore, we investigate the soft tissue interactions facilitated by the patient-side manipulator of the DaVinci surgical robot. Leveraging the pybullet physics engine, we simulate kinematics and establish anchor points to guide the robotic arm when manipulating soft tissue. Using demonstration-guided reinforcement learning (RL) algorithms, we investigate their performance in comparison to traditional reinforcement learning algorithms. Our in silico trials demonstrate a proof-of-concept for autonomous surgical soft tissue retraction. The results corroborate the feasibility of learning soft body manipulation through the application of reinforcement learning agents. This work lays the foundation for future research into the development and refinement of surgical robots capable of managing both rigid and soft tissue interactions. Code is available at https://github.com/amritpal-001/tissue_retract.
Improve Model Generalization and Robustness to Dataset Bias with Bias-regularized Learning and Domain-guided Augmentation
Nov 13, 2019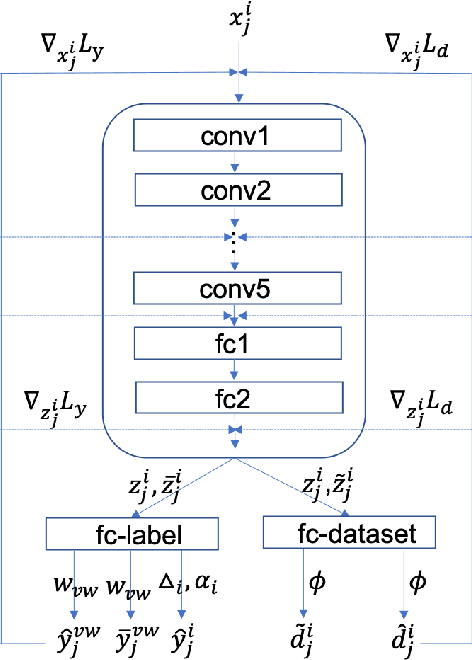
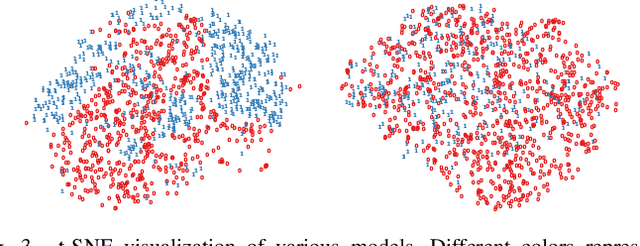
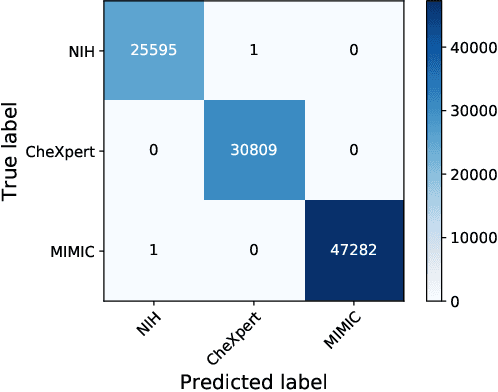

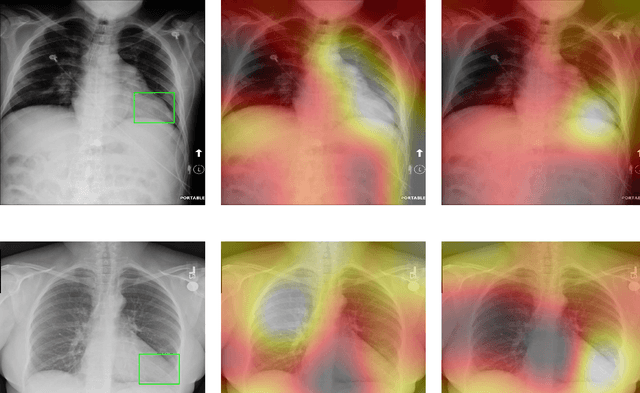
Abstract:Deep Learning has thrived on the emergence of biomedical big data. However, medical datasets acquired at different institutions have inherent bias caused by various confounding factors such as operation policies, machine protocols, treatment preference and etc. As the result, models trained on one dataset, regardless of volume, cannot be confidently utilized for the others. In this study, we investigated model robustness to dataset bias using three large-scale Chest X-ray datasets: first, we assessed the dataset bias using vanilla training baseline; second, we proposed a novel multi-source domain generalization model by (a) designing a new bias-regularized loss function; and (b) synthesizing new data for domain augmentation. We showed that our model significantly outperformed the baseline and other approaches on data from unseen domain in terms of accuracy and various bias measures, without retraining or finetuning. Our method is generally applicable to other biomedical data, providing new algorithms for training models robust to bias for big data analysis and applications. Demo training code is publicly available.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge