Martin Rektoris
Sparse Probabilistic Graph Circuits
Aug 11, 2025Abstract:Deep generative models (DGMs) for graphs achieve impressively high expressive power thanks to very efficient and scalable neural networks. However, these networks contain non-linearities that prevent analytical computation of many standard probabilistic inference queries, i.e., these DGMs are considered \emph{intractable}. While recently proposed Probabilistic Graph Circuits (PGCs) address this issue by enabling \emph{tractable} probabilistic inference, they operate on dense graph representations with $\mathcal{O}(n^2)$ complexity for graphs with $n$ nodes and \emph{$m$ edges}. To address this scalability issue, we introduce Sparse PGCs, a new class of tractable generative models that operate directly on sparse graph representation, reducing the complexity to $\mathcal{O}(n + m)$, which is particularly beneficial for $m \ll n^2$. In the context of de novo drug design, we empirically demonstrate that SPGCs retain exact inference capabilities, improve memory efficiency and inference speed, and match the performance of intractable DGMs in key metrics.
Probabilistic Graph Circuits: Deep Generative Models for Tractable Probabilistic Inference over Graphs
Mar 15, 2025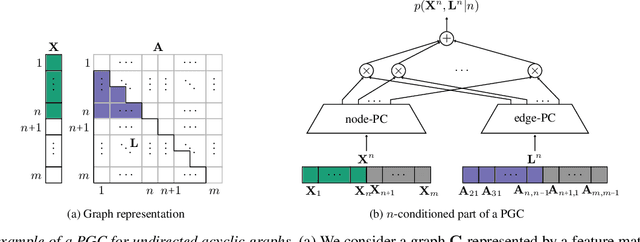
Abstract:Deep generative models (DGMs) have recently demonstrated remarkable success in capturing complex probability distributions over graphs. Although their excellent performance is attributed to powerful and scalable deep neural networks, it is, at the same time, exactly the presence of these highly non-linear transformations that makes DGMs intractable. Indeed, despite representing probability distributions, intractable DGMs deny probabilistic foundations by their inability to answer even the most basic inference queries without approximations or design choices specific to a very narrow range of queries. To address this limitation, we propose probabilistic graph circuits (PGCs), a framework of tractable DGMs that provide exact and efficient probabilistic inference over (arbitrary parts of) graphs. Nonetheless, achieving both exactness and efficiency is challenging in the permutation-invariant setting of graphs. We design PGCs that are inherently invariant and satisfy these two requirements, yet at the cost of low expressive power. Therefore, we investigate two alternative strategies to achieve the invariance: the first sacrifices the efficiency, and the second sacrifices the exactness. We demonstrate that ignoring the permutation invariance can have severe consequences in anomaly detection, and that the latter approach is competitive with, and sometimes better than, existing intractable DGMs in the context of molecular graph generation.
Sum-Product-Set Networks: Deep Tractable Models for Tree-Structured Graphs
Aug 18, 2024
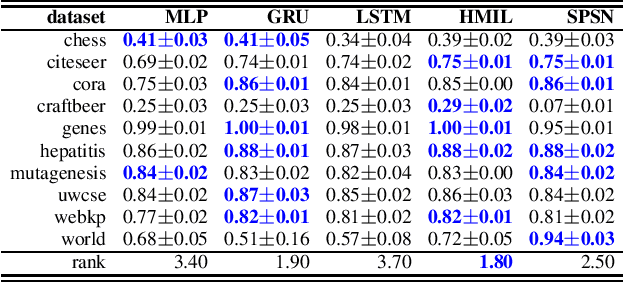

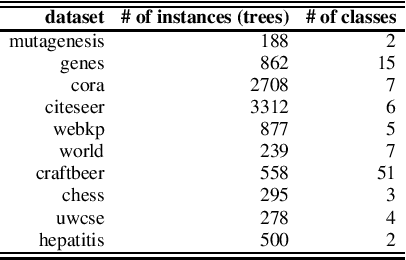
Abstract:Daily internet communication relies heavily on tree-structured graphs, embodied by popular data formats such as XML and JSON. However, many recent generative (probabilistic) models utilize neural networks to learn a probability distribution over undirected cyclic graphs. This assumption of a generic graph structure brings various computational challenges, and, more importantly, the presence of non-linearities in neural networks does not permit tractable probabilistic inference. We address these problems by proposing sum-product-set networks, an extension of probabilistic circuits from unstructured tensor data to tree-structured graph data. To this end, we use random finite sets to reflect a variable number of nodes and edges in the graph and to allow for exact and efficient inference. We demonstrate that our tractable model performs comparably to various intractable models based on neural networks.
GraphSPNs: Sum-Product Networks Benefit From Canonical Orderings
Aug 18, 2024Abstract:Deep generative models have recently made a remarkable progress in capturing complex probability distributions over graphs. However, they are intractable and thus unable to answer even the most basic probabilistic inference queries without resorting to approximations. Therefore, we propose graph sum-product networks (GraphSPNs), a tractable deep generative model which provides exact and efficient inference over (arbitrary parts of) graphs. We investigate different principles to make SPNs permutation invariant. We demonstrate that GraphSPNs are able to (conditionally) generate novel and chemically valid molecular graphs, being competitive to, and sometimes even better than, existing intractable models. We find out that (Graph)SPNs benefit from ensuring the permutation invariance via canonical ordering.
Sum-Product-Set Networks
Aug 14, 2024



Abstract:Daily internet communication relies heavily on tree-structured graphs, embodied by popular data formats such as XML and JSON. However, many recent generative (probabilistic) models utilize neural networks to learn a probability distribution over undirected cyclic graphs. This assumption of a generic graph structure brings various computational challenges, and, more importantly, the presence of non-linearities in neural networks does not permit tractable probabilistic inference. We address these problems by proposing sum-product-set networks, an extension of probabilistic circuits from unstructured tensor data to tree-structured graph data. To this end, we use random finite sets to reflect a variable number of nodes and edges in the graph and to allow for exact and efficient inference. We demonstrate that our tractable model performs comparably to various intractable models based on neural networks.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge