Marta B. M. Ranzini
MONAIfbs: MONAI-based fetal brain MRI deep learning segmentation
Mar 21, 2021
Abstract:In fetal Magnetic Resonance Imaging, Super Resolution Reconstruction (SRR) algorithms are becoming popular tools to obtain high-resolution 3D volume reconstructions from low-resolution stacks of 2D slices, acquired at different orientations. To be effective, these algorithms often require accurate segmentation of the region of interest, such as the fetal brain in suspected pathological cases. In the case of Spina Bifida, Ebner, Wang et al. (NeuroImage, 2020) combined their SRR algorithm with a 2-step segmentation pipeline (2D localisation followed by a 2D segmentation network). However, if the localisation step fails, the second network is not able to recover a correct brain mask, thus requiring manual corrections for an effective SRR. In this work, we aim at improving the fetal brain segmentation for SRR in Spina Bifida. We hypothesise that a well-trained single-step UNet can achieve accurate performance, avoiding the need of a 2-step approach. We propose a new tool for fetal brain segmentation called MONAIfbs, which takes advantage of the Medical Open Network for Artificial Intelligence (MONAI) framework. Our network is based on the dynamic UNet (dynUNet), an adaptation of the nnU-Net framework. When compared to the original 2-step approach proposed in Ebner-Wang, and the same Ebner-Wang approach retrained with the expanded dataset available for this work, the dynUNet showed to achieve higher performance using a single step only. It also showed to reduce the number of outliers, as only 28 stacks obtained Dice score less than 0.9, compared to 68 for Ebner-Wang and 53 Ebner-Wang expanded. The proposed dynUNet model thus provides an improvement of the state-of-the-art fetal brain segmentation techniques, reducing the need for manual correction in automated SRR pipelines. Our code and our trained model are made publicly available at https://github.com/gift-surg/MONAIfbs.
Combining multimodal information for Metal Artefact Reduction: An unsupervised deep learning framework
Apr 20, 2020
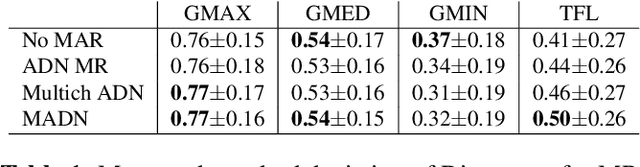

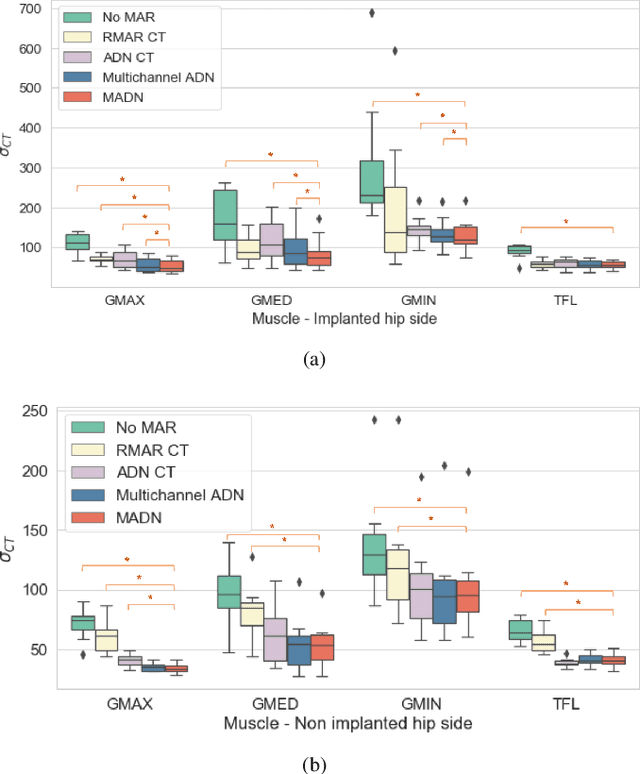
Abstract:Metal artefact reduction (MAR) techniques aim at removing metal-induced noise from clinical images. In Computed Tomography (CT), supervised deep learning approaches have been shown effective but limited in generalisability, as they mostly rely on synthetic data. In Magnetic Resonance Imaging (MRI) instead, no method has yet been introduced to correct the susceptibility artefact, still present even in MAR-specific acquisitions. In this work, we hypothesise that a multimodal approach to MAR would improve both CT and MRI. Given their different artefact appearance, their complementary information can compensate for the corrupted signal in either modality. We thus propose an unsupervised deep learning method for multimodal MAR. We introduce the use of Locally Normalised Cross Correlation as a loss term to encourage the fusion of multimodal information. Experiments show that our approach favours a smoother correction in the CT, while promoting signal recovery in the MRI.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge