Maria Leptin
Stateless actor-critic for instance segmentation with high-level priors
Jul 06, 2021

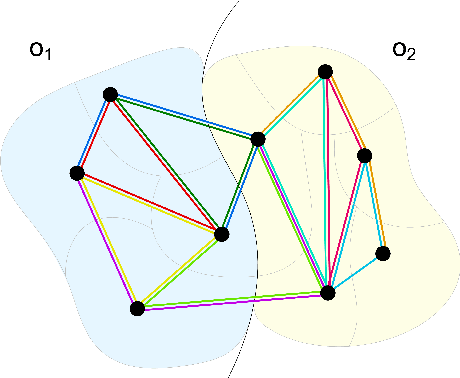
Abstract:Instance segmentation is an important computer vision problem which remains challenging despite impressive recent advances due to deep learning-based methods. Given sufficient training data, fully supervised methods can yield excellent performance, but annotation of ground-truth data remains a major bottleneck, especially for biomedical applications where it has to be performed by domain experts. The amount of labels required can be drastically reduced by using rules derived from prior knowledge to guide the segmentation. However, these rules are in general not differentiable and thus cannot be used with existing methods. Here, we relax this requirement by using stateless actor critic reinforcement learning, which enables non-differentiable rewards. We formulate the instance segmentation problem as graph partitioning and the actor critic predicts the edge weights driven by the rewards, which are based on the conformity of segmented instances to high-level priors on object shape, position or size. The experiments on toy and real datasets demonstrate that we can achieve excellent performance without any direct supervision based only on a rich set of priors.
Semi-Automatic Generation of Tight Binary Masks and Non-Convex Isosurfaces for Quantitative Analysis of 3D Biological Samples
Jan 30, 2020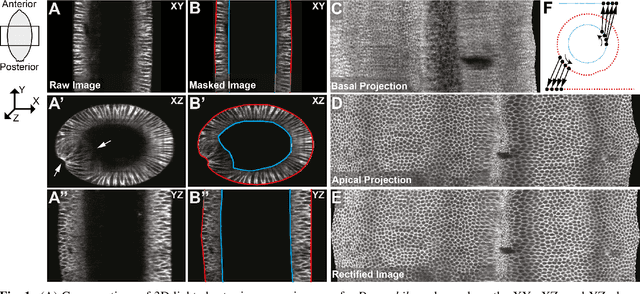

Abstract:Current in vivo microscopy allows us detailed spatiotemporal imaging (3D+t) of complete organisms and offers insights into their development on the cellular level. Even though the imaging speed and quality is steadily improving, fully-automated segmentation and analysis methods are often not accurate enough. This is particularly true while imaging large samples (100um - 1mm) and deep inside the specimen. Drosophila embryogenesis, widely used as a developmental paradigm, presents an example for such a challenge, especially where cell outlines need to imaged - a general challenge in other systems as well. To deal with the current bottleneck in analyzing quantitatively the 3D+t light-sheet microscopy images of Drosophila embryos, we developed a collection of semi-automatic open-source tools. The presented methods include a semi-automatic masking procedure, automatic projection of non-convex 3D isosurfaces to 2D representations as well as cell segmentation and tracking.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge