Maksim Sharaev
Integrating Radiomics with Deep Learning Enhances Multiple Sclerosis Lesion Delineation
Jun 17, 2025



Abstract:Background: Accurate lesion segmentation is critical for multiple sclerosis (MS) diagnosis, yet current deep learning approaches face robustness challenges. Aim: This study improves MS lesion segmentation by combining data fusion and deep learning techniques. Materials and Methods: We suggested novel radiomic features (concentration rate and R\'enyi entropy) to characterize different MS lesion types and fused these with raw imaging data. The study integrated radiomic features with imaging data through a ResNeXt-UNet architecture and attention-augmented U-Net architecture. Our approach was evaluated on scans from 46 patients (1102 slices), comparing performance before and after data fusion. Results: The radiomics-enhanced ResNeXt-UNet demonstrated high segmentation accuracy, achieving significant improvements in precision and sensitivity over the MRI-only baseline and a Dice score of 0.774$\pm$0.05; p<0.001 according to Bonferroni-adjusted Wilcoxon signed-rank tests. The radiomics-enhanced attention-augmented U-Net model showed a greater model stability evidenced by reduced performance variability (SDD = 0.18 $\pm$ 0.09 vs. 0.21 $\pm$ 0.06; p=0.03) and smoother validation curves with radiomics integration. Conclusion: These results validate our hypothesis that fusing radiomics with raw imaging data boosts segmentation performance and stability in state-of-the-art models.
Weakly Supervised Fine Tuning Approach for Brain Tumor Segmentation Problem
Nov 06, 2019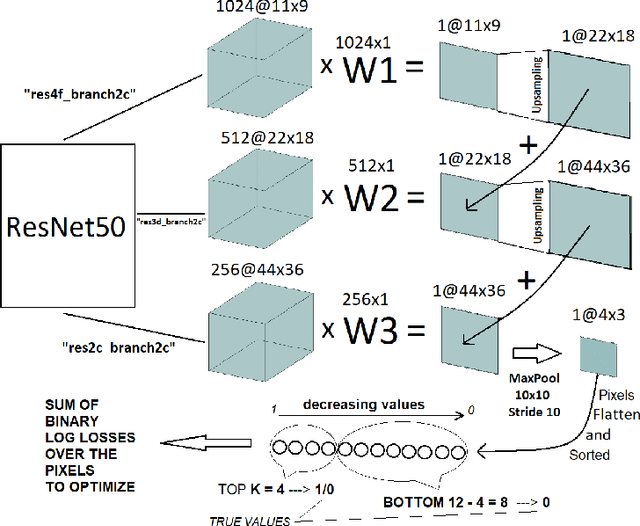
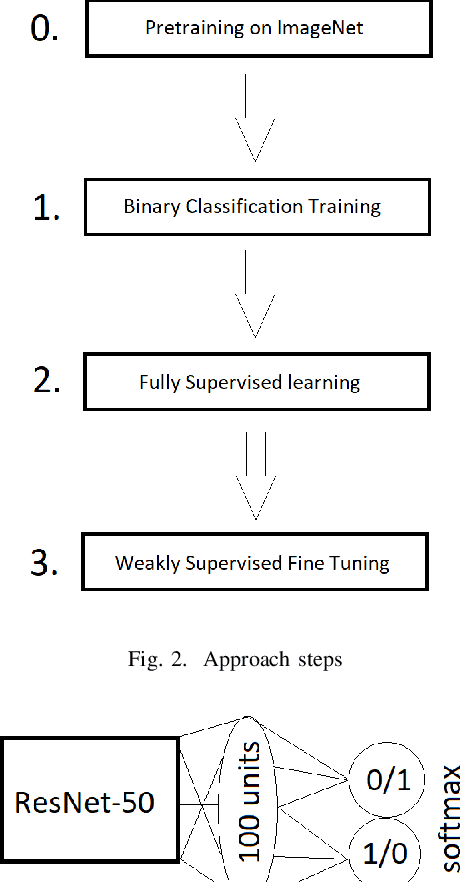
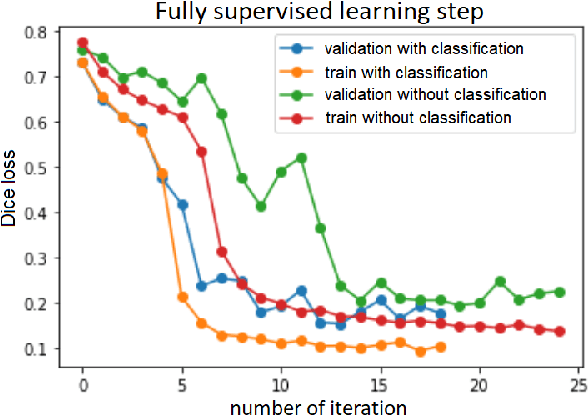
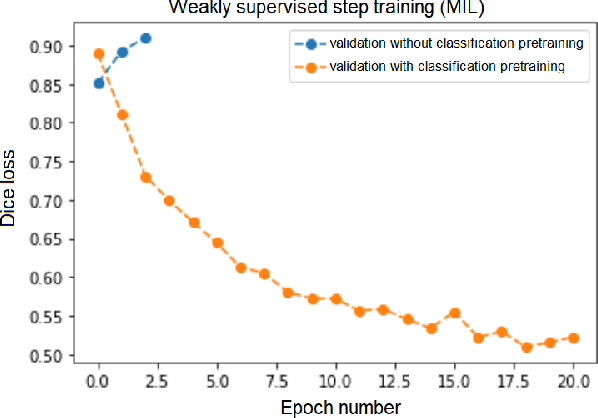
Abstract:Segmentation of tumors in brain MRI images is a challenging task, where most recent methods demand large volumes of data with pixel-level annotations, which are generally costly to obtain. In contrast, image-level annotations, where only the presence of lesion is marked, are generally cheap, generated in far larger volumes compared to pixel-level labels, and contain less labeling noise. In the context of brain tumor segmentation, both pixel-level and image-level annotations are commonly available; thus, a natural question arises whether a segmentation procedure could take advantage of both. In the present work we: 1) propose a learning-based framework that allows simultaneous usage of both pixel- and image-level annotations in MRI images to learn a segmentation model for brain tumor; 2) study the influence of comparative amounts of pixel- and image-level annotations on the quality of brain tumor segmentation; 3) compare our approach to the traditional fully-supervised approach and show that the performance of our method in terms of segmentation quality may be competitive.
3D Deformable Convolutions for MRI classification
Nov 05, 2019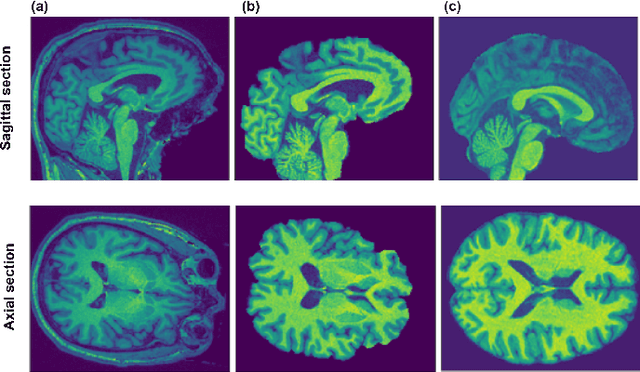
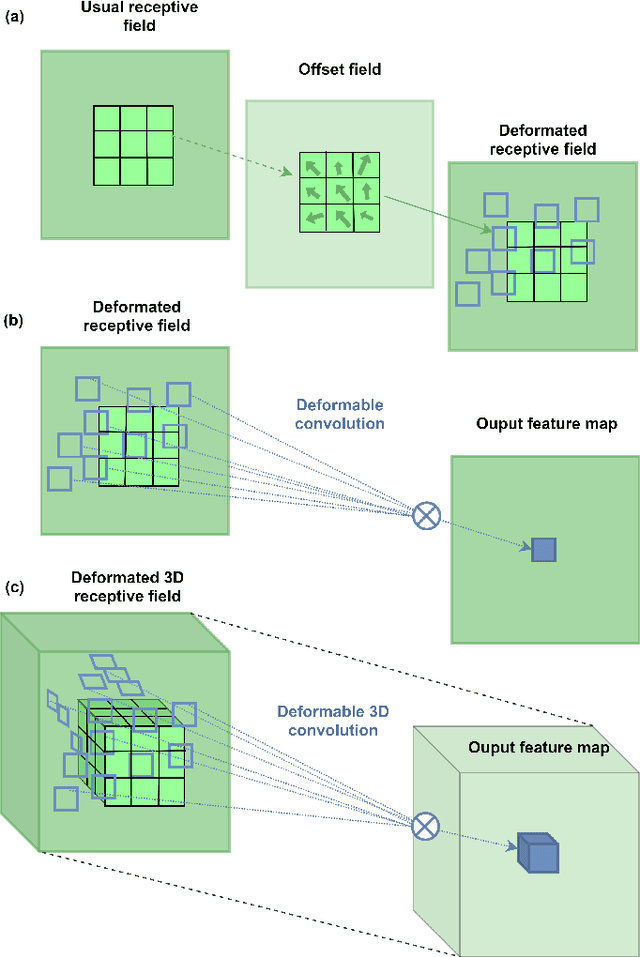
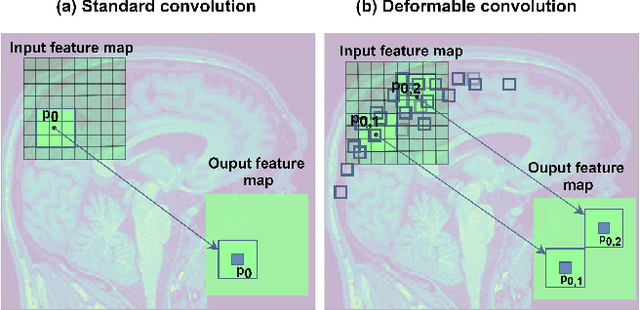
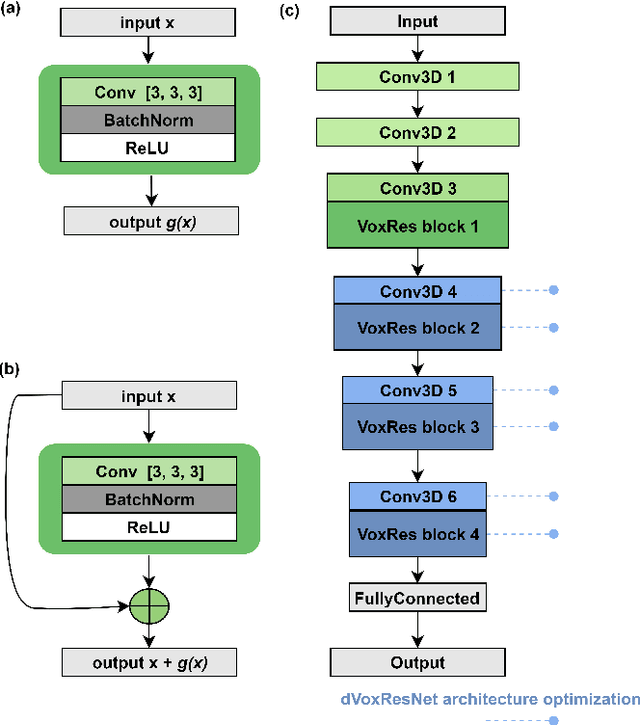
Abstract:Deep learning convolutional neural networks have proved to be a powerful tool for MRI analysis. In current work, we explore the potential of the deformable convolutional deep neural network layers for MRI data classification. We propose new 3D deformable convolutions(d-convolutions), implement them in VoxResNet architecture and apply for structural MRI data classification. We show that 3D d-convolutions outperform standard ones and are effective for unprocessed 3D MR images being robust to particular geometrical properties of the data. Firstly proposed dVoxResNet architecture exhibits high potential for the use in MRI data classification.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge